Raman pixel by pixel
<br>
Raman spectroscopy provides molecular specificity through spectrally-resolved measurement of the inelastic scattering under monochromatic excitation. In the context of microscopy, it may serve as label-free cell imaging, providing structural information.
However, the very low cross-section of Raman scattering requires long time exposures, which preclude imaging of cellular components with low concentrations. Surface-enhanced Raman spectroscopy (SERS), which relies on the local electromagnetic field enhancement produced by metallic nanostructures, is an approach to drastically increase the sensitivity of the Raman detection while retaining large amounts of spectral information.
In cellular imaging, the measurement is usually performed on endocytosed nanostructures. However, the measured SERS signals vary strongly as they depend on excitation beam profile, local particle presence or aggregation and local molecular environment. Identifying and extracting spectra corresponding to molecules of interest within a SERS data set is very difficult.
Conventional data analysis methods look for global patterns in the data, whereas the singlemolecule sensitivity of SERS can detect independent molecules in each pixel with little correlation between pixels. Nicolas Pavillon and his colleagues from Osaka University now explored different algorithmic methods to automatically discriminate spectra of interest in the measured field of view, without imposing assumptions on the self-similarity of the data.
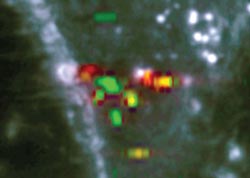
The proposed method relies on the indexing of the positions of relevant spectra, which are selected by the computation of a quality map. The scientists proposed various criteria to compute spectra extraction, such as the spectral energy, the peak count per spectra, or the projection coefficients on SVD vectors. They assessed each criteria with simulated data and applied this approach to different types of measurements, such as dried Rhodamine 6G adsorbed on gold nanoparticles deposited on a glass substrate, and HeLa cells with endocytosed gold nanoparticles.
The tests with simulated data showed that various criteria can provide satisfactory results. The computation time could be tremendously decreased by discarding irrelevant pixels through a simple criterion based on the spectral energy, reducing the processing time to typically less than 10 seconds for a field of view on the order of 100 X 100 pixels. The tests performed on Rhodamine 6G measurements demonstrated the validity of the proposed approach, where its known spectrum could be extracted automatically.
The peak count criterion was the most suitable for most cases, as it detects various patterns without filtering out any curve which may only appear a single instance in the data set. Such single spectra may be critical important in a given SERS detection experiment. One main feature of the proposed approach is that its output is a localization map of the most relevant spectra in a measurement. The spatial information is retained, making it possible to trace back the positions of several spectra with identical properties, for instance. The optimized method was utilized to extract and classify the complex SERS response behavior of gold nanoparticles taken in live cells. (Text contributed by K. Maedefessel-Herrmann)
N. Pavillon, K. Bando, K. Fujita, N. I. Smith, Feature-based recognition of Surface-enhanced Raman spectra for biological targets, J. Biophotonics 6(8),587-597 (2013); doi: http://dx.doi.org/10.1002/jbio.201200181
For more information about the Journal of Biophotonics visit the journal homepage.
Regina Hagen
Journal Publishing Manager, Journal of Biophotonics
Managing Editor, Physical Sciences
Global Research
Wiley-VCH Verlag GmbH & Co. KGaA
Rotherstrasse 21
10245 Berlin
Germany
www.wiley.com
T +49 (0)30 47 031 321
F +49 (0)30 47 031 399
jbp@wiley.com
www.biophotonics-journal.org
www.wileyonlinelibrary.com
Media Contact
More Information:
http://www.wiley.comAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















