Biologists Find ‘Missing Link’ in the Production of Protein Factories in Cells
Their discovery, detailed in the June 23 issue of the journal Genes & Development, will not only force a revision of basic textbooks on molecular biology, but also provide scientists with a better understanding of how to limit uncontrolled cell growth, such as cancer, that might be regulated by controlling the output of ribosomes.
Ribosomes are responsible for the production of the wide variety of proteins that include enzymes; structural molecules, such as hair, skin and bones; hormones like insulin; and components of our immune system such as antibodies.
Regarded as life’s most important molecular machine, ribosomes have been intensively studied by scientists (the 2009 Nobel Prize in Chemistry, for example, was awarded for studies of its structure and function). But until now researchers had not uncovered all of the details of how the proteins that are used to construct ribosomes are themselves produced.
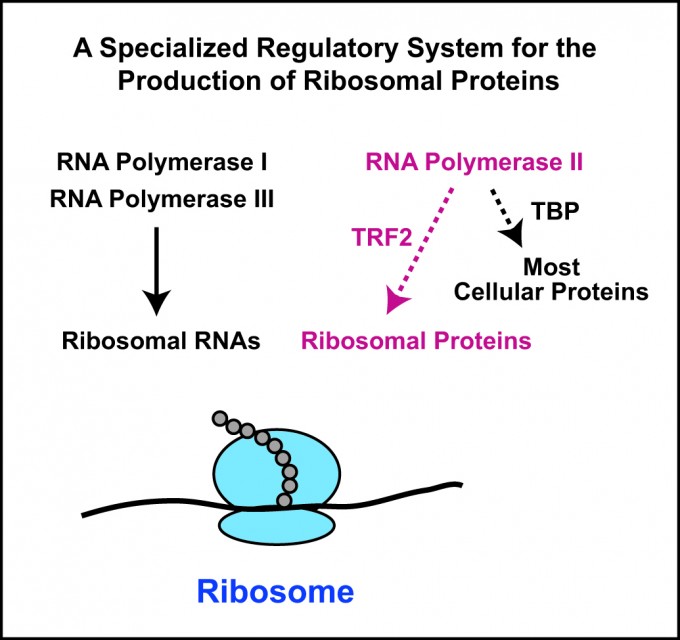
In multicellular animals such as humans, ribosomes are made up of about 80 different proteins (humans have 79 while some other animals have a slightly different number) as well as four different kinds of RNA molecules. In 1969, scientists discovered that the synthesis of the ribosomal RNAs is carried out by specialized systems using two key enzymes: RNA polymerase I and RNA polymerase III. But until now, scientists were unsure if a complementary system was also responsible for the production of the 80 proteins that make up the ribosome.
That’s essentially what the UC San Diego researchers headed by Jim Kadonaga, a professor of biology, set out to examine. What they found was the missing link—the specialized system that allows ribosomal proteins themselves to be synthesized by the cell.
“We found that ribosomal proteins are synthesized via a novel regulatory system with the enzyme RNA polymerase II and a factor termed TRF2,” Kadonaga says. “For the production of most proteins, RNA polymerase II functions with a factor termed TBP, but for the synthesis of ribosomal proteins, it uses TRF2.”
“The discovery of this specialized TRF2-based system for ribosome biogenesis,” he adds, “provides a new avenue for the study of ribosomes and its control of cell growth, and should lead to a better understanding and potential treatment of diseases such as cancer.”
Other authors of the paper were UC San Diego biologists Yuan-Liang Wang, Sascha Duttke and George Kassavetis, and Kai Chen, Jeff Johnston, and Julia Zeitlinger of the Stowers Institute for Medical Research in Kansas City, Missouri. Their research was supported by two grants from the National Institutes of Health (1DP2OD004561-01 and R01 GM041249).
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















