New technique puts DNA profiling of E. coli on fast track

Michigan State University has developed a new technique to test the DNA of E. coli bacteria by examining very small genetic changes called single nucleotide polymorphisms or SNPs (pronounced snips). Using SNPs, scientists analyzed 96 markers, making genetic analysis of pathogenic bacteria possible at a rate never before accomplished.
“It used to take three months to score one gene individually,” said Thomas Whittam, Hannah Distinguished Professor at the National Food Safety and Toxicology Center at MSU. “Now, we are working on a new, more rapid system that can do thousands of genes per day.”
In a new study released in the Monday edition of the Proceedings of the National Academy of Sciences, “Variation in Virulence Among Clades of Escherichia coli O157:H7 Associated With Disease Outbreaks,” Whittam and his co-authors looked at the DNA of more than 500 strains of a particularly dangerous member of the E. coli family, O157:H7. In collaboration with David Alland of the University of Medicine and Dentistry of New Jersey, Whittam discovered that individual bacteria could be separated into nine major groups, called clades.
E coli makes people sick because they produce toxins, called Shiga toxins. These toxins block protein synthesis, an essential cellular function, particularly in the kidneys. What Whittam found was that the different clades produced different kinds of Shiga toxins in varying amounts based on their DNA.
“For the first time, we know why some outbreaks cause serious infections and diseases and others don’t,” Whittam said. “The different E. coli groups produce different toxins.”
Rapid genetic characterization also opens up a new world of possibilities for identifying the bacterial culprits in outbreaks and finding out where they originated.
E. coli usually come from animal waste contaminating human sources of food or water. Finding out how the bacteria entered the food source always has been a challenge, but now food safety experts can use DNA just like police use DNA at crime scenes. Scientists will be able to identify those bacteria making people sick, find out where they entered the food source and then use this information to reduce contamination.
“This is the first time anyone has been able to classify very closely related groups,” Whittam said.
“This is also the first time we can tell the differences in how they cause disease.”
Whittam also has plans to use this methodology to study other bacterial strains, like Shigella, a major cause of diarrhea around the world. “This new equipment can be used to identify hundreds of thousands of pathogenic bacteria,” Whittam said.
Media Contact
More Information:
http://www.msu.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
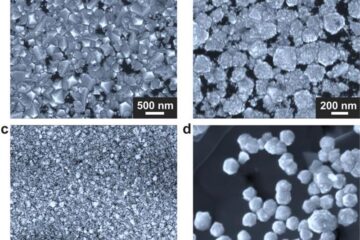

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…





















