Software helps decrypt embryonic development
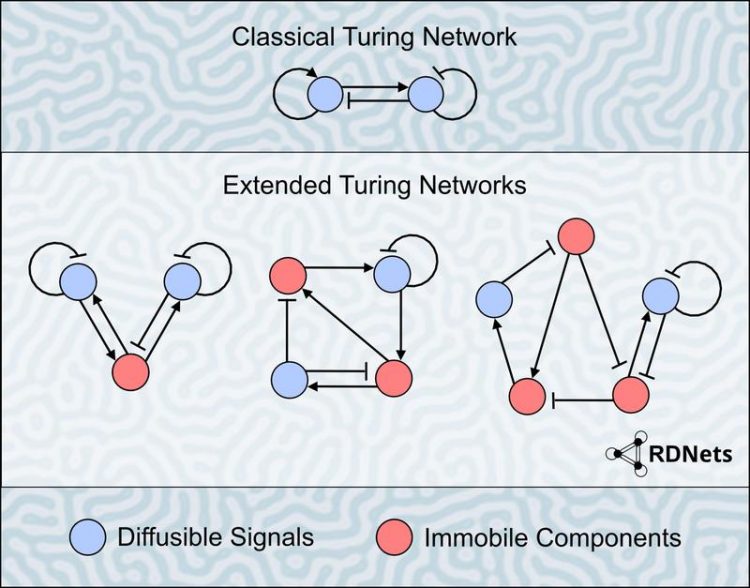
Model of a classical Turing network compared to the extended Turing networks analyzed with the software RDNets. Background: a self-organizing labyrinthine Turing pattern Luciano Marcon/ Friedrich Miescher Laboratory Tübingen
More than six decades ago, Alan Turing – the father of modern computer science, famous for decrypting messages from the Enigma machine during World War II – postulated a model explaining how the body is patterned during embryonic development. In the reaction-diffusion model that he proposed, two signaling molecules react with each other and spread through the embryo by diffusion.
Turing showed mathematically that the two molecules can form spatial patterns in the embryo if one molecule moves faster than the other. The resulting high and low molecule concentrations could then provide the cells with information on how to differentiate and where to form different body parts.
Although Turing’s patterns remarkably resemble the patterns observed during normal development, Turing models have been limited to two mobile signaling molecules and could not take into account our current knowledge about the complex underlying gene regulatory networks, which have only been identified over the last few decades after Turing’s death.
Now, a paper published in the journal eLife extends Turing’s original approach to the post-genomic era. To include the effect that gene regulatory networks have on pattern formation, a team led by Dr. Patrick Müller with scientists from the Friedrich Miescher Laboratory in Tübingen and the Centre for Genomic Regulation in Barcelona developed a new computational method to analyze and simulate the formation of patterns in reaction-diffusion networks with both mobile and immobile molecules.
“Real-life patterning systems don´t consist of the simplified two-component networks used in classical Turing models. We wanted to analyze more complex systems and to create a user-friendly software that allows us to uncover new biologically relevant network designs”, explains Dr. Luciano Marcon, first author of the study.
The analysis of Turing models involves tedious mathematics, and it takes approximately two pages to analyze a given network by hand. Extending this approach to a systematic analysis with millions of possible networks would seem impossible. The scientists therefore used a modern computer algebra system and developed the software RDNets, which can perform the tedious mathematics automatically within a few minutes.
Screening millions of possible reaction-diffusion networks with their software, the scientists discovered that most of the newly identified patterning networks do not need to fulfill the condition of differential signal mobility that Turing postulated and that has been thought to be indispensable. Instead, patterns can also form when the signaling molecules are equally mobile or even with any combination of signal mobilities. “We have found that realistic reaction-diffusion systems follow mechanisms that are fundamentally different from the previous concepts”, says Dr. Patrick Müller.
The scientists used their software to analyze several developmental systems, from the generation of progenitor tissues to the formation of fingers. In addition to its relevance for developmental biology, RDNets may also be useful for bioengineers. The software enables users to model many patterning processes and to design underlying gene regulatory circuits that can then be built synthetically. This should be of great use for tissue engineering approaches, where the software user can design regions of interest for the expression of differentiating factors to induce specific tissues in defined domains.
About the software
Systems biology focuses on the study of biochemical networks, where molecules are the nodes and the molecular interactions are the edges. The dynamics of these networks can be described by differential equations that represent the behavior of the molecules in space and time. There are two ways to study differential equations: either by finding an analytical solution to the equations, or by performing numerical simulations. Traditionally, the first approach is performed manually by mathematicians who use algebra to write down a closed-form solution. This solution describes the behavior of the system for all possible parameter values. The second approach is instead performed by computers and involves the execution of thousands of repetitive calculations with the aim to obtain a list of numerical values that describe the behavior of the system under specific conditions, i.e. for a set of representative parameter values. Therefore, analytical approaches are preferable because they provide an exhaustive description of the system, in contrast to numerical simulations which can only sample a subset of the parameter space. In practice, however, analytical approaches are limited to simple networks because the mathematics becomes too complicated as the size of the problem grows, and the manual solution of the equations becomes unfeasible.
The software RDNets published in eLife was able to overcome this limitation by automating a linear stability analysis of partial differential equations with the aid of a computer algebra system. This novel approach of an automated high-throughput mathematical analysis can be used to screen for new biochemical Turing networks that can form self-organizing periodic patterns. The analysis can be constrained with qualitative and quantitative experimental data, which makes RDNets an unprecedented tool for users that aim to study developmental patterning networks or to design reaction-diffusion synthetic circuits. The software is available online at http://www.RDNets.com and runs on most web browsers.
Original Publication
Marcon L, Diego X, Sharpe J, Müller P. High-throughput mathematical analysis identifies Turing networks for patterning with equally diffusing signals. eLife. 2016 Apr 8;5. pii: e14022. doi: 10.7554/eLife.14022.
Contact:
Dr. Patrick Müller
Max Planck Research Group “Systems Biology of Development”
phone: 07071 601-815
email: patrick.mueller@tuebingen.mpg.de
http://www.fml.tuebingen.mpg.de/mueller-group.html
Nadja Winter (PR Officer)
phone: +49 7071 601- 444
email: presse-eb@tuebingen.mpg.de
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

A universal framework for spatial biology
SpatialData is a freely accessible tool to unify and integrate data from different omics technologies accounting for spatial information, which can provide holistic insights into health and disease. Biological processes…

How complex biological processes arise
A $20 million grant from the U.S. National Science Foundation (NSF) will support the establishment and operation of the National Synthesis Center for Emergence in the Molecular and Cellular Sciences (NCEMS) at…

Airborne single-photon lidar system achieves high-resolution 3D imaging
Compact, low-power system opens doors for photon-efficient drone and satellite-based environmental monitoring and mapping. Researchers have developed a compact and lightweight single-photon airborne lidar system that can acquire high-resolution 3D…





















