Precise, Highly Efficient Gene Repair
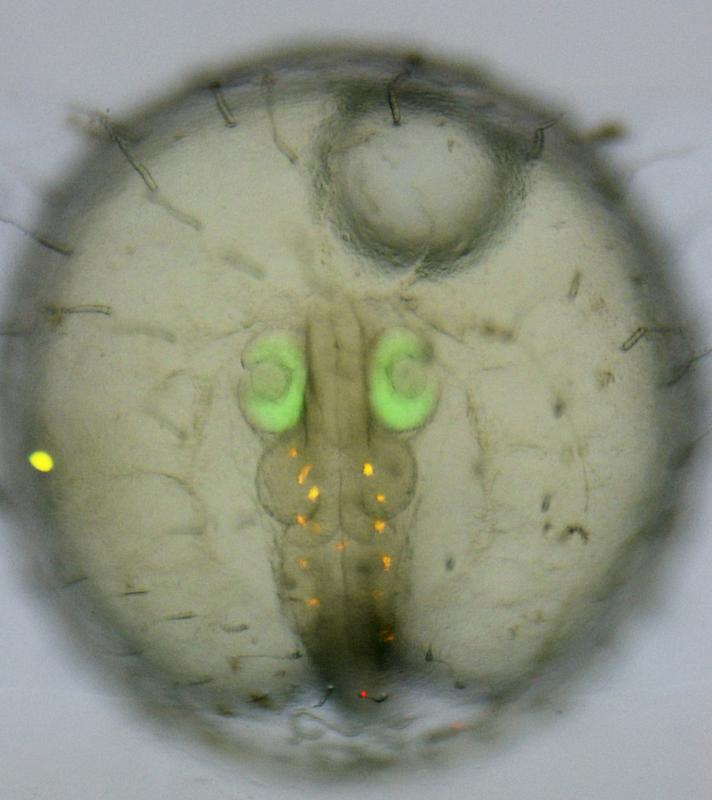
Medaka embryo, in which the Rx2 gene – a gene critical for eye development – was edited with a repair copy. (Complete caption: see text Gutierrez-Triana, Tavhelidse, Thumberger et al., 2018, figure three, subject to CC BY 4.0 license
The molecular tool CRISPR/Cas allows introducing DNA double strand breaks into any gene of interest consequently resulting in stochastic mutations at the site of the target gene. However, precise gene repair through the application of a rescue construct suffers from limited efficiency.
Researchers at Heidelberg University have now found a solution for this problem. Applying their new approach on the model organism medaka, the researchers laid the groundwork for easily integrating the repair copy of a defective gene into the DNA.
As developmental biologist Prof. Dr Joachim Wittbrodt explains, this efficient process makes precise genome editing possible in basic research, bringing the tool much closer to its application in medical treatment. The research results were published in “eLife”.
Gene editing involves finding and targeting the precise location in the genome that causes gene mutations. A guide RNA is constructed for this task. It consists of RNA sequences that are complementary to the target DNA sequence.
The guide RNA docks at the desired location of the DNA where the CRISPR/Cas as a pair of molecular “scissors” cuts the double strand. The cell's own repair system then jumps into action. Individual DNA building blocks can get lost upon repair of these breaks, causing stochastic mutations in selected target genes.
According to Prof. Wittbrodt of Heidelberg University‘s Centre for Organismal Studies, this method has been used successfully in virtually every organism studied.
However, routine use of precise editing that result in exactly defined modifications in any gene remained elusive so far. As Dr Arturo Gutierrez explains, the cell's own repair system, which quickly closes the breaks in the double DNA strand, is to blame. Whereas this mechanism, known as “Non-Homologous End Joining” (NHEJ), is not a problem for efficiently introducing stochastic genetic modifications, it competes with a second, extremely precise repair process called “Homology Directed Repair” (HDR).
Just like when a replacement part is installed, both ends have to fit perfectly, so that HDR can replace the defective gene with the correct repair copy. “Unfortunately, NHEJ connects the copies introduced into the cells into long continuous chains, rendering them unusable,” says Tinatini Tavhelidse.
Using the Japanese ricefish medaka, Prof. Wittbrodt's team developed and validated a new approach that enables highly efficient, precise genetic repair, which is a basic prerequisite for application in gene surgery.
The Heidelberg researchers pursued a simple idea. Instead of using pharmacological compounds with serious side effects to mitigate the unwanted effects of NHEJ, they modified the repair copy such that it could not be “attacked” and rendered useless. Both ends of the copy are blocked by biotin – a B vitamin – to prevent non-homologous end joining. “This inexpensive process now enables efficient gene repair by precisely inserting a single repair copy,” adds Dr Thomas Thumberger.
Caption:
Medaka embryo, in which the Rx2 gene – a gene critical for eye development – was edited with a repair copy. This copy contains the sequence for a green fluorescent protein.
Source: Gutierrez-Triana, Tavhelidse, Thumberger et al., 2018, figure three, subject to CC BY 4.0 license
Contact:
Communications and Marketing
Press Office, phone +49 6221 54-2311
presse@rektorat.uni-heidelberg.de
Prof. Dr Joachim Wittbrodt
Centre for Organismal Studies
Phone +49 6221 54-6499
jochen.wittbrodt@cos.uni-heidelberg.de
J.A. Gutierrez-Triana, T. Tavhelidse, T. Thumberger, I. Thomas, B. Wittbrodt, T. Kellner, K. Anlas, E. Tsingos and J. Wittbrodt: Efficient single-copy HDR by 5’ modified long dsDNA donors. eLife, https://doi.org/10.7554/eLife.39468
https://www.cos.uni-heidelberg.de/index.php/index.php/j.wittbrodt?l=_e
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















