Mechanism of an enzyme for biofuel production
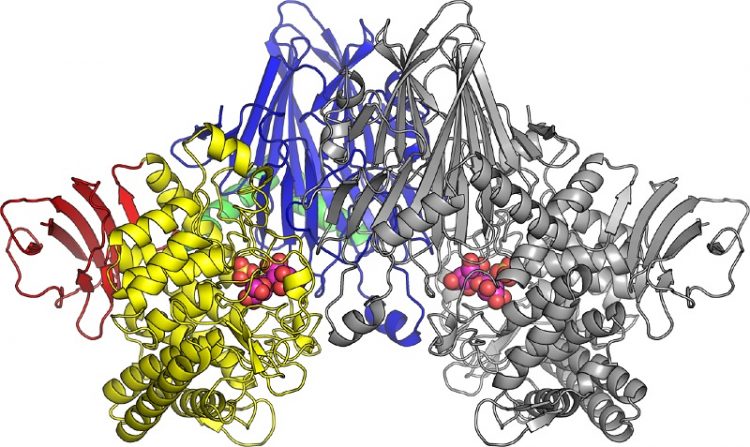
*The three-dimensional structure of cellobionic acid phopshorylase. © 2015 Shinya Fushinobu.
A University of Tokyo research group has revealed for the first time the three-dimensional structure and mechanism of action of a key enzyme of bio-fuel production, cellobionic acid phosphorylase (CBAP). This result is important basic information for developing the technology to make bio-fuel and other chemical products from biomass.
It has been long thought that hydrolytic enzymes (cellulases) were the main contributors to microbial degradation of cellulose. Recently, the existence of oxidative cellulose-degrading enzymes that dramatically increase the activity efficiency of cellulases have been noted.
When these enzymes degrade cellulose, cellobionic acid is produced. However, it was completely unknown how the cellulolytic microbes further metabolize this compound.
In 2013, one of the members of the research group, Associate Professor Hiroyuki Nakai at the Graduate School of Science and Technology, Niigata University, discovered a new enzyme, cellobionic acid phosphorylase (CBAP).
CBAP catalyzes the degradation of cellobionic acid to produce compounds that are prone to further metabolism and fermentation. Therefore, this enzyme is a missing link between the oxidative cellulose degradation and bioethanol fermentation pathways in microorganisms. However, the three dimensional structure of the enzyme and the mechanism by which it degraded cellobionic acid remained unknown.
In this latest research, the research group lead by Professor Shinya Fushinobu at the University of Tokyo, Graduate School of Agricultural and Life Sciences, used X-ray crystallography to reveal the three-dimensional structure of CBAP isolated from marine bacteria. In addition, the structure of CBAP in complex with cellobionic acid was determined (figure), and the reaction mechanism for decomposing cellobionic acid was revealed.
“This research is extremely interesting from a scientific perspective, but could also contribute to the development of biorefinery technologies that produce biofuels such as ethanol and other useful compounds via biomass degradation by microbes,” says Professor Fushinobu.
*Image
Two cellobionic acid phosphorylases molecules pair up to create a dimer. The colored left half and the gray right half are each one enzyme molecule. Cellobionic acid and sulfuric acid ions (a compound similar to phosphoric acid) bound to cellobionic acid phosphorylase are expressed as spheres in the figure.
Paper
Young-Woo Nam, Takanori Nihira, Takatoshi Arakawa, Yuka Saito, Motomitsu Kitaoka, Hiroyuki Nakai, and Shinya Fushinobu, “Crystal structure and substrate recognition of cellobionic acid phosphorylase playing a key role in oxidative cellulose degradation by microbes”, The Journal of Biological Chemistry Vol. 290, No. 30, pg 18281-18292, doi: 10.1074/jbc.M115.664664.
Associated links
UTokyo Research article
Media Contact
More Information:
http://www.researchsea.comAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















