Genome tree of life is largest yet for seed plants
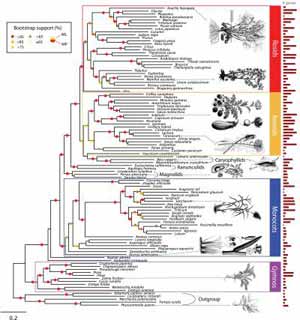
This is a phylogenomic reconstruction of the evolutionary diversification of seed plants. Credit: E.K. Lee et al.<br>
Scientists at the American Museum of Natural History, Cold Spring Harbor Laboratory, The New York Botanical Garden, and New York University have created the largest genome-based tree of life for seed plants to date.
Their findings, published today in the journal PLoS Genetics, plot the evolutionary relationships of 150 different species of plants based on advanced genome-wide analysis of gene structure and function. This new approach, called “functional phylogenomics,” allows scientists to reconstruct the pattern of events that led to the vast number of plant species and could help identify genes used to improve seed quality for agriculture.
“Ever since Darwin first described the 'abominable mystery' behind the rapid explosion of flowering plants in the fossil record, evolutionary biologists have been trying to understand the genetic and genomic basis of the astounding diversity of plant species,” said Rob DeSalle, a corresponding author on the paper and a curator in the Museum's Division of Invertebrate Zoology who conducts research at the Sackler Institute for Comparative Genomics. “Having the architecture of this plant tree of life allows us to start to decipher some of the interesting aspects of evolutionary innovations that have occurred in this group.”
The research, performed by members of the New York Plant Genomics Consortium, was funded by the National Science Foundation (NSF) Plant Genome Program to identify the genes that caused the evolution of seeds, a trait of important economic interest. The group selected 150 representative species from all of the major seed plant groups to include in the study. The species span from the flowering variety—peanuts and dandelions, for example—to non-flowering cone plants like spruce and pine. The sequences of the plants' genomes—all of the biological information needed to build and maintain an organism, encoded in DNA—were either culled from pre-existing databases or generated, in the field and at The New York Botanical Garden in the Bronx, from live specimens.
With new algorithms developed at the Museum and NYU and the processing power of supercomputers at Cold Spring Harbor Laboratory and overseas, the sequences—nearly 23,000 sets of genes (specific sections of DNA that code for certain proteins)—were grouped, ordered, and organized in a tree according to their evolutionary relationships. Algorithms that determine similarities of biological processes were used to identify the genes underlying species diversity.
“Previously, phylogenetic trees were constructed from standard sets of genes and were used to identify the relationships of species,” said Gloria Coruzzi, a professor in New York University's Center for Genomics and Systems Biology and the principal investigator of the NSF grant. “In our novel approach, we create the phylogeny based on all the genes in a genome, and then use the phylogeny to identify which genes provide positive support for the divergence of species.”
The results support major hypotheses about evolutionary relationships in seed plants. The most interesting finding is that gnetophytes, a group that consists mostly of shrubs and woody vines, are the most primitive living non-flowering seed plants—present since the late Mesozoic era, the “age of dinosaurs.” They are situated at the base of the evolutionary tree of seed plants.
“This study resolves the long-standing problem of producing an unequivocal evolutionary tree of the seed plants,” said Dennis Stevenson, vice president for laboratory research at The New York Botanical Garden. “We can then use this information to determine when and where important adaptations occur and how they relate to plant diversification. We also can examine the evolution of such features as drought tolerance, disease resistance, or crop yields that sustain human life through improved agriculture.”
In addition, the researchers were able to make predictions about genes that caused the evolution of important plant characteristics. One such evolutionary signal is RNA interference, a process that cells use to turn down or silence the activity of specific genes. Based on their new phylogenomic maps, the researchers believe that RNA interference played a large role in the separation of monocots—plants that have a single seed leaf, including orchids, rice, and sugar cane—from other flowering plants. Even more surprising, RNA interference also played a major role in the emergence of flowering plants themselves.
“Genes required for the production of small RNA in seeds were at the very top of the list of genes responsible for the evolution of flowering plants from cone plants,” said Rob Martienssen, a professor at Cold Spring Harbor Laboratory. “In collaboration with colleagues from LANGEBIO [Laboratorio Nacional de Genomica para la Biodiversidad] in Mexico last year, we found that these same genes control maternal reproduction, providing remarkable insight into the evolution of reproductive strategy in flowering plants.”
The data and software resources generated by the researchers are publicly available and will allow other comparative genomic researchers to exploit plant diversity to identify genes associated with a trait of interest or agronomic value. These studies could have implications for improving the quality of seeds and, in turn, agricultural products ranging from food to clothing.
In addition, the phylogenomic approach used in this study could be applied to other groups of organisms to further explore how species originated, expanded, and diversified.
“The collaboration among the institutions involved here is a great example of how modern science works,” said Sergios-Orestis Kolokotronis, a term assistant professor at Columbia University's Barnard College and a research associate at the Museum's Sackler Institute. “Each of the four institutions involved has its own strengths and these strengths were nicely interwoven to produce a novel vision of plant evolution.”
Other authors include Ernest Lee, American Museum of Natural History; Angelica Cibrian-Jaramillo, American Museum of Natural History, The New York Botanical Garden, and New York University – currently at the Laboratorio Nacional de Genomica para la Biodiversidad, Mexico; Manpreet Katari, New York University; Alexandros Stamatakis, Technical University Munich – currently at Heidelberg Institute for Theoretical Studies; Michael Ott, Technical University Munich; Joanna Chiu, University of California, Davis; Damon Little, The New York Botanical Garden; and W. Richard McCombie, Cold Spring Harbor Laboratory.
Media Contact
More Information:
http://www.amnh.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















