An embryonic cell’s fate is sealed by the speed of a signal
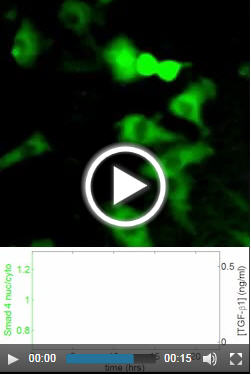
Tempting fate. To visualize cells’ responses to the signals that ultimately lead them to choose a fate, the researchers engineered a protein involved in this response, Smad4, to glow. In response to a pulse of signal molecules, Smad4 moves into the dark nuclei of the cells, causing them to glow briefly.
“It turns out that if ramped up slowly enough an otherwise potent signal elicits no response from the receiving cells. Meanwhile, a pulsing, on-off signal appears to have a stronger effect than a constant one,” says researcher Ali Brivanlou, Robert and Harriet Heilbrunn Professor and head of the Laboratory of Molecular Vertebrate Embryology. This research is the latest collaboration between Brivanlou and Eric Siggia, Viola Ward Brinning and Elbert Calhoun Brinning Professor at Rockefeller’s Center for Studies in Physics and Biology.
“Until now, it has not been feasible to test how speed or other temporal dynamics affect a cell’s response to a signal. However, by adapting technology that allows for very precise control over these aspects, we found unequivocal evidence that signal level alone does not determine a cell’s fate. Its presentation is also extremely important,” Siggia says.
Together, the team dubbed their discovery “speed fating.” Their work will be published in August in Developmental Cell.
Biologists know a cell determines its location in an embryo and, as a result, its future role, based on chemical cues from its neighbors. About 50 years ago, the developmental biologist Lewis Wolpert proposed that this determination hinges on the concentration of the signal to which a cell is exposed: Go above a certain threshold and you get one fate, below and you get a second. His proposal is known as the French flag model, after a tri-color graph used to represent three cell fates based on those cells’ positions with respect to the source of the signal.
Prior work from Brivanlou and Siggia had cast doubt on the sole importance of concentration. Using a common developmental signaling pathway known as TGF-β, the team documented what is known as an adaptive response from cells exposed to TGF-β signaling molecules. This response peaked then declined over time, even though the signaling molecules remained present. (Think of how a constant noise eventually blends into the background.) If concentration was the sole factor responsible for a response, then the response should have continued as long as the signal was present.
To follow up on this work, Benoit Sorre, a former Rockefeller postdoc now at the University of Paris Diderot, adapted a system that makes use of miniaturized networks of pipes, pumps, valves and sample chambers all under computer control. For experiments, he teamed up with Aryeh Warmflash, the postdoc who lead the previous TGF-β work. Together, they worked with mouse cells that have the potential to differentiate into muscle, or cartilage and bone. Progenitor cells like these, which can differentiate into a limited set of tissues, are the offspring of stem cells. In experiments using Sorre’s new system, the researchers exposed these progenitor cells to signaling molecules from the TGF-β pathway, and then recorded the cells’ responses to see if the signal activated the pathway that leads them to choose a fate.
Sorre and Warmflash started with a continuous signal. As Warmflash’s previous work suggested, this finger-stuck-on-the-buzzer approach did not produce a continuous response from the cells. Instead, the response declined. A second set of tests showed a series of brief pulses of signal produced a greater response than one continuous signal.
Gradually increasing the concentration of the signal, however, appeared to have the opposite effect. The researchers ramped up the concentration of the signal over periods as brief as five hours or as long as 40 hours. The longer the period and the slower the rate of increase, the weaker the cells’ response. The cells subjected to a 40-hour run barely registered at all.
Based on these experiments, the team formulated a mathematical model to describe how a cell in an embryo may infer its position in relation to the source of the signal. In this way, the research offers a new take on the French flag model: It is still true that the fates of three cells can be mapped out based on their position, but the cells appear to arrive at these fates more rapidly than previously thought, thanks to the adaptive response that takes into account both the level and speed of a signal.
“This finding is another instance of a productive collaboration between biologists and physicists. Neither group, biologists or physicists, could have realized this result working alone,” Siggia says.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

A universal framework for spatial biology
SpatialData is a freely accessible tool to unify and integrate data from different omics technologies accounting for spatial information, which can provide holistic insights into health and disease. Biological processes…

How complex biological processes arise
A $20 million grant from the U.S. National Science Foundation (NSF) will support the establishment and operation of the National Synthesis Center for Emergence in the Molecular and Cellular Sciences (NCEMS) at…

Airborne single-photon lidar system achieves high-resolution 3D imaging
Compact, low-power system opens doors for photon-efficient drone and satellite-based environmental monitoring and mapping. Researchers have developed a compact and lightweight single-photon airborne lidar system that can acquire high-resolution 3D…





















