A social network of human proteins
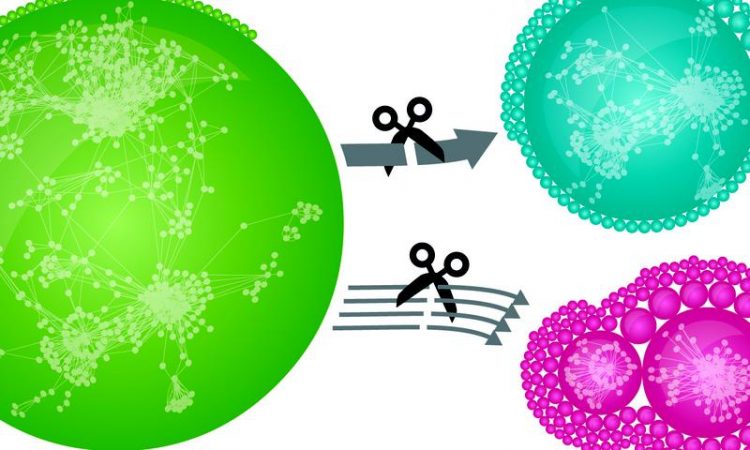
Thousands of human proteins are connected via tens of thousands of distinct interactions in a network (green). If weak interactions are removed, defined local modules emerge (red). Removal of strong interactions has a much smaller effect on network structure (blue). This underlines the relevance of weak interactions for global interconnectedness. © MPI of Biochemistry
Scientists at the Max Planck Institute of Biochemistry in Martinsried near Munich and at the MPI of Molecular Cell Biology and Genetics in Dresden have now drawn a detailed map of human protein interactions.
Using a novel mass spectrometric quantification method, the researchers determined the strength of each interaction. “Our data revealed that most interactions are weak, but critical for the structure of the entire network,” explains Marco Hein, first author of the study. The paper has now been published in the Journal Cell.
Proteins are the building blocks and central protagonists of the cell and contribute to all processes of life at the molecular level. They carry out their tasks by binding to each other and building interaction networks. With the help of quantitative mass spectrometry, scientists can determine precisely which proteins interact with each other.
The technology can be described as molecular fishing: One protein is selected as bait. Fishing it out of a complex mixture retrieves all its interaction partners as well, which are then identified by a mass spectrometer. Scientists from Martinsried and Dresden have now analyzed 1,100 such bait proteins in a large-scale project. They mapped a network of over 5,400 proteins, which are connected by 28,000 interactions.
The different interactions have very distinct properties. Some connections are strong and serve a structural role, others are weak and transient, for instance in signal transduction pathways. Measuring the strength of an interaction is very laborious and hence complicated in high throughput studies.
Using a novel strategy, the German scientists established a method of estimating the strength of each interaction indirectly. They measure the copy numbers of all proteins in the cell, and quantify the ratio at which each interactor is retrieved along with its corresponding bait protein. The stronger an interaction, the more of an interactor is recovered.
The study offers a new perspective of the “social network” of human proteins. Researchers can now not only browse the new repository to see which proteins interact with each other. They can also distinguish different types of interactions.
The data reveal that weak interactions dominate the network. “A single weak interaction may seem irrelevant. In their entirety, however, these interactions form the backbone that holds the entire network together,” explains Marco Hein. “This is a property that the network of proteins has in common with the social network in society.”
Contact
Dr. Christiane Menzfeld
Max Planck Institute of Biochemistry, Martinsried
Phone: +49 89 8578-2824
Fax: +49 89 8578-3777
Email: menzfeld@biochem.mpg.de
Prof. Dr. Matthias Mann
Max Planck Institute of Biochemistry, Martinsried
Phone: +49 89 8578-2557
Fax: +49 89 8578-2219
Email: mmann@biochem.mpg.de
Original publication
M.Y. Hein, N.C. Hubner, I. Poser, J. Cox, N. Nagaraj, Y. Toyoda, I.A. Gak, I. Weisswange, J. Mansfeld, F. Buchholz, A.A. Hyman & M. Mann:
A human interactome in three quantitative dimensions organized by stoichiometries and abundances.
Cell, October, 2015 DOI: 10.1016/j.cell.2015.09.053
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

A universal framework for spatial biology
SpatialData is a freely accessible tool to unify and integrate data from different omics technologies accounting for spatial information, which can provide holistic insights into health and disease. Biological processes…

How complex biological processes arise
A $20 million grant from the U.S. National Science Foundation (NSF) will support the establishment and operation of the National Synthesis Center for Emergence in the Molecular and Cellular Sciences (NCEMS) at…

Airborne single-photon lidar system achieves high-resolution 3D imaging
Compact, low-power system opens doors for photon-efficient drone and satellite-based environmental monitoring and mapping. Researchers have developed a compact and lightweight single-photon airborne lidar system that can acquire high-resolution 3D…





















