Researchers demonstrate new technique that improves the power of atomic force micrscopy
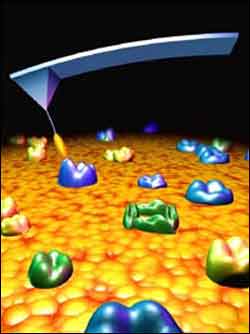
An artist’s depiction shows an atomic force microscope probe (not to scale) ’fishing’ for molecular sites recognized by an antibody tethered to the probe by a fine polymer thread. The new technique promises to vastly improve the capabilities of atomic force microscopy.
A team of researchers have developed a method that could vastly improve the ability of atomic force microscopes to “see” the chemical composition of a sample, follow variations of the sample, as well as map its topographic structure.
The advance could have significant implications for drug development by allowing scientists to monitor the effects of potential drugs on an ever-smaller scale, according to Stuart Lindsay, director of the Center for Single Molecule Biophysics at the Biodesign Institute at Arizona State University and a lead researcher on the project.
Lindsay, an ASU professor in the department of physics and astronomy said the new technique allows an atomic force microscope to “see,” on a nanometer scale, the chemical composition of molecules.
“Atomic force microscopy has a resolution down to an atomic level, but until now it has been blind to identifying specific chemical compositions,” Lindsay said.
The researchers — Lindsay, Hongda Wang, Ralph Bash, Brian Ashcroft, and Dennis Lohr of Arizona State University; Cordula Stroh, Hermann Gruber and Peter Hinterdorfer of the Institute of Biophysics at the University of Lintz, Austria; and Jeremy Nelson of Molecular Imaging Corporation, Tempe, Ariz. — present their findings in “Single Molecule Recognition Imaging Microscopy” in the current issue of the Proceedings of the National Academy of Sciences. The article is available on line at http://www4.nationalacademies.org/nas/nashome.nsf
“If you imagine that all proteins are shaped like Lego blocks, then conventional atomic force microscopy (AFM) is feeling the Lego blocks on the floor, but it can’t tell the difference between one block and another,” Lindsay explained. “What we have done, is allow the person sitting on the floor and feeling those blocks to open their eyes and see that there are red Lego blocks, green Lego blocks and yellow Lego blocks.”
“This allows you to identify specific components in an image,” he added. “It means you can now follow a complex process and see what’s happening, at the molecular level, to one of the components. We are now giving AFM chemical sensitivity in much the way colored dyes gave optical microscopes optical sensitivity for much larger objects (~1 micron).”
Atomic force microscopes provide images on the nanometer scale by using a highly sensitive and tiny probe that is essentially pulled across a surface. By doing this, researchers can obtain topographical images down to a nanometer scale.
To use the AFM in its new mode, the researchers attached antibodies keyed to individual proteins to the tip of an AFM’s probe. When an antibody reacts with the protein it is specifically targeted for, it creates a variance in the microscope’s reading compared to a reading with a bare tip, thus showing the presence of a protein or other specific material in the region being scanned.
To help ensure that the antibody tipped probe is truly sensitive, a strand of polymer connects the antibody to the tip, providing a tether that allows the antibody to wiggle into position to better connect with the protein receptors. A magnetically excited cantilever makes the tip oscillate up and down to make the antibody disconnect and reconnect and keep the probe moving.
A key capability of this technique, Lindsay said, is that it allows researchers to see how components of a cell react on a molecular scale when they experience biological processes, such as their response to a specific chemical or compound. In this mode, it could provide researchers with a molecular “time-lapsed movie” of such reactions, which could lead to greater understanding of the chemical dynamics involved in how cells react to such stimuli.
Lindsay said the new AFM method could be significant for drug discovery.
“This development opens up the AFM as a research tool,” Lindsay added. “The ability to identify the specific proteins on a membrane surface means you can take something very complex, like the surface of a human cell with all of the types of different receptors on it and ask questions about the local chemistry, like what is binding at those sites. That will provide the fundamental knowledge you need to develop new drugs.”
Media Contact
More Information:
http://www.asu.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…





















