Indispensable Guests: Shedding Light on Past Life
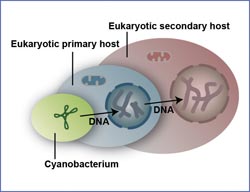
Friendly takeover: In secondary endosymbiosis, a bacteria (green) wanders into the primary host (blue), which then absorbs a secondary host (red); the DNA is also carried over in the process.<br><br>(Image: John M. Archibald, Dalhousie University, Canada)<br>
The hosts haven’t transferred all of their permanent guests’ genetic material into their nuclei, because genetic information could have become lost in the process, as the scientists assume. Their results appear in the current issue of the journal Nature from Thursday, 6 December 2012.
“For the first time, we have sequenced the nuclear genomes of two unicellular algae that are remarkable in their genetic and cellular complexity,” write the authors, among whom are Professor Dr. Stefan Rensing, Professor Dr. Uwe Maier, Dr. Franziska Hempel, Aikaterini Symeonidi, and Dr. Stefan Zauner from the University of Marburg. Cells contain a variety of so-called organelles – subcellular components that fulfill critical functions for the cell, such as energy conversion or photosynthesis. These organelles originated from previously discrete cells that were integrated into the cell in the distant past by the ancestors of the host organism.
Usually organelles were converted from bacteria, but not so in the case of several species of algae: They learned photosynthesis by absorbing other plant cells, including all of their existing organelles, the so-called chloroplasts. The term for this process is secondary endosymbiosis: By continuing to take advantage of the services of their new workers, the hosts can use sunlight to produce nutrients.
Structures that do not serve this purpose are usually lost in the course of evolution. In a few rare cases, however, the symbiotic organelles still contain nuclei of greatly reduced dimensions that stem from the original organism – for instance in cryptophytes and chlorarachniophytes.
Why does the nucleus remain intact in these species? In order to find out, the international research consortium first determined which sequences the four nuclear genomes of two relevant species possess. The result: Both of the residual nuclei contained only a fraction of the genes possessed by independently living related species. As the authors report, much of the remaining genetic information does not exhibit any similarity at all to that of known genes. Nevertheless, they control vitally important processes that occur in these organelles, such as translation, i.e., the conversion of genetic information into proteins.
A sizable quantity of genes evidently wandered from the organelles acquired by way of secondary endosymbiosis into the nucleus of the host. The scientists refer to the result as a “a complex mosaic of genes whose evolutionary histories do not reliably predict where their protein products function within the cell.”
Uwe Maier is a member of “LOEWE – Center for Synthetic Microbiology” of the University of Marburg. Among other things, he and his team in Marburg prepared sequencing data for the project. In addition, they identified encoded proteins and determined their location. Stefan Rensing’s research group at the Cluster of Excellence “BIOSS Centre for Biological Signalling Studies” of the University of Freiburg helped to identify the proteins that are imported into the organelles and analyzed contaminations, duplication events, and all of the proteins involved in the regulation of the transcription.
(Translation: University of Freiburg)
Original Publication: Bruce A. Curtis & al.: Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs, Nature (6.12.2012), DOI: 10.1038/nature11681
Further Information:
Professor Dr. Stefan Rensing,
Fachgebiet Zellbiologie
E-Mail: stefan.rensing@biologie.uni-marburg.de
Professor Dr. Uwe Maier,
Fachgebiet Zellbiologie und „LOEWE-Zentrum für Synthetische Mikrobiologie (SYNMIKRO)“
Tel.: 06421/28-21543
E-Mail: maier@biologie.uni-marburg.de
Media Contact
More Information:
http://www.uni-marburg.deAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

A universal framework for spatial biology
SpatialData is a freely accessible tool to unify and integrate data from different omics technologies accounting for spatial information, which can provide holistic insights into health and disease. Biological processes…

How complex biological processes arise
A $20 million grant from the U.S. National Science Foundation (NSF) will support the establishment and operation of the National Synthesis Center for Emergence in the Molecular and Cellular Sciences (NCEMS) at…

Airborne single-photon lidar system achieves high-resolution 3D imaging
Compact, low-power system opens doors for photon-efficient drone and satellite-based environmental monitoring and mapping. Researchers have developed a compact and lightweight single-photon airborne lidar system that can acquire high-resolution 3D…





















