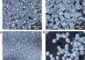
Nanoribbons in solutions mimic nature
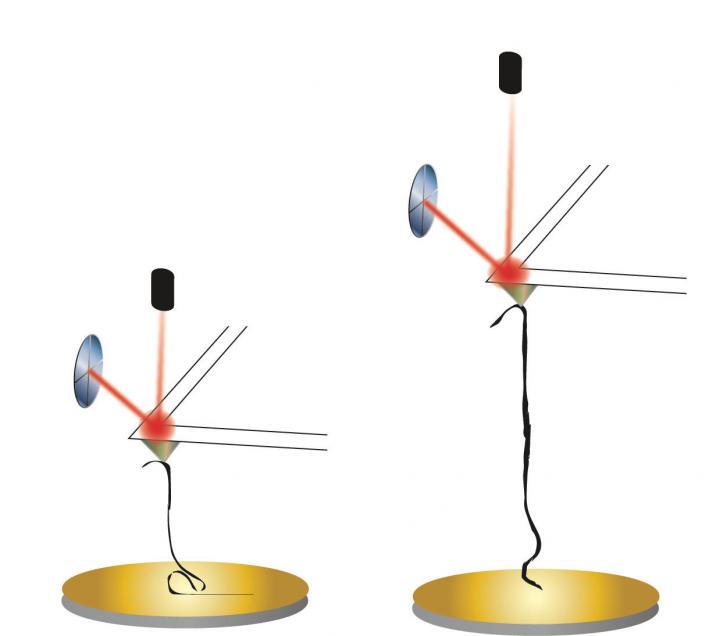
The tip of an atomic force microscope on a cantilevered arm is used to pull a graphene nanoribbon the same way it would be used to pull apart a protein or a strand of DNA in a Rice University lab. The microscope can be used to measure properties like rigidity in a material as it's manipulated by the tip. Credit: Kiang Research Group/Rice University
Graphene nanoribbons (GNRs) bend and twist easily in solution, making them adaptable for biological uses like DNA analysis, drug delivery and biomimetic applications, according to scientists at Rice University.
Knowing the details of how GNRs behave in a solution will help make them suitable for wide use in biomimetics, according to Rice physicist Ching-Hwa Kiang, whose lab employed its unique capabilities to probe nanoscale materials like cells and proteins in wet environments. Biomimetic materials are those that imitate the forms and properties of natural materials.
The research led by recent Rice graduate Sithara Wijeratne, now a postdoctoral researcher at Harvard University, appears in the Nature journal Scientific Reports.
Graphene nanoribbons can be thousands of times longer than they are wide. They can be produced in bulk by chemically “unzipping” carbon nanotubes, a process invented by Rice chemist and co-author James Tour and his lab.
Their size means they can operate on the scale of biological components like proteins and DNA, Kiang said. “We study the mechanical properties of all different kinds of materials, from proteins to cells, but a little different from the way other people do,” she said. “We like to see how materials behave in solution, because that's where biological things are.” Kiang is a pioneer in developing methods to probe the energy states of proteins as they fold and unfold.
She said Tour suggested her lab have a look at the mechanical properties of GNRs. “It's a little extra work to study these things in solution rather than dry, but that's our specialty,” she said.
Nanoribbons are known for adding strength but not weight to solid-state composites, like bicycle frames and tennis rackets, and forming an electrically active matrix. A recent Rice project infused them into an efficient de-icer coating for aircraft.
But in a squishier environment, their ability to conform to surfaces, carry current and strengthen composites could also be valuable.
“It turns out that graphene behaves reasonably well, somewhat similar to other biological materials. But the interesting part is that it behaves differently in a solution than it does in air,” she said. The researchers found that like DNA and proteins, nanoribbons in solution naturally form folds and loops, but can also form helicoids, wrinkles and spirals.
Kiang, Wijeratne and Jingqiang Li, a co-author and student in the Kiang lab, used atomic force microscopy to test their properties. Atomic force microscopy can not only gather high-resolution images but also take sensitive force measurements of nanomaterials by pulling on them. The researchers probed GNRs and their precursors, graphene oxide nanoribbons.
The researchers discovered that all nanoribbons become rigid under stress, but their rigidity increases as oxide molecules are removed to turn graphene oxide nanoribbons into GNRs. They suggested this ability to tune their rigidity should help with the design and fabrication of GNR-biomimetic interfaces.
“Graphene and graphene oxide materials can be functionalized (or modified) to integrate with various biological systems, such as DNA, protein and even cells,” Kiang said. “These have been realized in biological devices, biomolecule detection and molecular medicine. The sensitivity of graphene bio-devices can be improved by using narrow graphene materials like nanoribbons.”
Wijeratne noted graphene nanoribbons are already being tested for use in DNA sequencing, in which strands of DNA are pulled through a nanopore in an electrified material. The base components of DNA affect the electric field, which can be read to identify the bases.
The researchers saw nanoribbons' biocompatibility as potentially useful for sensors that could travel through the body and report on what they find, not unlike the Tour lab's nanoreporters that retrieve information from oil wells.
Further studies will focus on the effect of the nanoribbons' width, which range from 10 to 100 nanometers, on their properties.
###
Co-authors are Rice research scientist Evgeni Penev; graduate student Wei Lu; alumna Amanda Duque, now a scientist at Los Alamos National Laboratory; and Boris Yakobson, the Karl F. Hasselmann Professor of Materials Science and NanoEngineering and a professor of chemistry. Tour is the T.T. and W.F. Chao Professor of Chemistry as well as a professor of computer science and of materials science and nanoengineering. Kiang is an associate professor of physics and astronomy and of bioengineering.
The Welch Foundation and the National Science Foundation supported the research. The researchers used the NSF's Extreme Science and Engineering Discovery Environment and the NSF-supported DAVinCI supercomputer administered by Rice's Center for Research Computing and procured in a partnership with Rice's Ken Kennedy Institute.
Read the abstract at http://www.
This news release can be found online at http://news.
Follow Rice News and Media Relations via Twitter @RiceUNews
Related materials:
Ching-Hwa Kiang Research Group: http://www.
Wiess School of Natural Sciences: http://natsci.
Images for download:
http://news.
Rice University physicist Ching-Hwa Kiang, left, and graduate student Jingqiang Li analyze readings at the atomic force microscope in her lab. The researchers analyzed the properties of carbon nanoribbons in solutions with the equipment they normally use to analyze DNA, proteins and cells. (Credit: Rice University)
http://news.
Rice University physicist Ching-Hwa Kiang, left, and alumna Sithara Wijeratne led a study to determine the mechanical properties of graphene nanoribbons in solution. They found the nanoribbons mimic the properties of natural polymers like proteins and DNA and may be suitable for biomimetic applications. (Credit: Jeff Fitlow/Rice University)
http://news.
The tip of an atomic force microscope on a cantilevered arm is used to pull a graphene nanoribbon the same way it would be used to pull apart a protein or a strand of DNA in a Rice University lab. The microscope can be used to measure properties like rigidity in a material as it's manipulated by the tip. (Credit: Kiang Research Group/Rice University)
Located on a 300-acre forested campus in Houston, Rice University is consistently ranked among the nation's top 20 universities by U.S. News & World Report. Rice has highly respected schools of Architecture, Business, Continuing Studies, Engineering, Humanities, Music, Natural Sciences and Social Sciences and is home to the Baker Institute for Public Policy. With 3,910 undergraduates and 2,809 graduate students, Rice's undergraduate student-to-faculty ratio is 6-to-1. Its residential college system builds close-knit communities and lifelong friendships, just one reason why Rice is ranked No. 1 for best quality of life and for lots of race/class interaction by the Princeton Review. Rice is also rated as a best value among private universities by Kiplinger's Personal Finance. To read “What they're saying about Rice,” go to http://tinyurl.
Media Contact
All latest news from the category: Materials Sciences
Materials management deals with the research, development, manufacturing and processing of raw and industrial materials. Key aspects here are biological and medical issues, which play an increasingly important role in this field.
innovations-report offers in-depth articles related to the development and application of materials and the structure and properties of new materials.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















