Search for macular degeneration genes narrows

Scientists zero in on five chromosome regions
Scientists at the University of Michigan Kellogg Eye Center, working with colleagues in the U-M School of Public Health, have significantly narrowed the range of chromosomal locations where they expect to find genes associated with age-related macular degeneration (AMD).
In a paper published in the March issue of American Journal of Human Genetics, Kellogg scientist Anand Swaroop, Ph.D., and his team of researchers have confirmed three previously suggested loci (on chromosomes 1, 5, and 9) for potential AMD genes, and have identified two new loci on chromosomes 2 and 22.
AMD is a progressive disease that destroys central vision. There is no known cure for the disease, which affects millions of individuals worldwide. While scientists believe there is a strong genetic component, most believe the cause will be found in the interplay of several genes combined with environmental factors, such as smoking and diet.
Swaroop says that this study is significant because it confirms locations that have been suggested by other researchers, noting that cross-validation is important for complex diseases such as AMD.
“Our researchers have been able to build on past studies — our own and a handful of others — and now we are ready to begin the search for AMD susceptibility genes in earnest,” says Swaroop, a professor of ophthalmology and of human genetics at the U-M Medical School. “We have identified very specific regions of the genome where we can intensify our research efforts.”
Kellogg researchers narrowed the search for AMD-related genes by performing a high-resolution genome scan of all 23 pairs of participants’ chromosomes, using over 700 DNA markers. Markers are known DNA segments that help define locations and regions on chromosomes. The markers were placed approximately five mega-bases apart (on average), thereby delimiting the regions on chromosomes where researchers can continue their search for susceptibility genes.
Patients were recruited from the Kellogg Eye Center’s clinical practice and consisted largely of individuals who suffer from late-stage AMD (geographic atrophy or choroidal neovascularization). Swaroop observes that this group is likely to carry a heavier load of genetic susceptibility factors than patients in earlier stages of the disease, and thus would be well suited for genetic investigations.
In addition, each of the affected individuals was paired with a sibling or other relative (total of 412 affected relative pairs), creating a greater likelihood of studying subjects with similar genetic profiles.
The Kellogg research team could not replicate the finding of an earlier study indicating that a mutation in the gene Hemicentitin-1 might be linked with AMD. After studying over 800 individuals, the Kellogg team saw no evidence of the variant, but suggested that another gene in the same region on chromosome 1 may undergo a change that predisposes individuals to AMD.
The next step for Kellogg researchers is to use microarray-based methods to identify candidate genes within the newly defined regions, says Swaroop. “We will further narrow the candidate regions using other genetic methods, in addition to performing association studies with candidate genes. The next few years should be very exciting,” he adds.
Swaroop, who collaborated on this study with Goncolo R. Abecasis, D.Phil., U-M School of Public Health, notes that Kellogg is well positioned to conduct genetic investigations because collaboration between clinicians and researchers is encouraged and convenient.
Kellogg’s Retina Service clinicians, in particular, made important contributions to the study. In addition, Kellogg’s Center for Retinal and Macular Degeneration, under Swaroop’s direction, has one of the largest patient resources for AMD genetic studies, with 2,114 individuals representing 1,534 families currently enrolled.
The paper is titled “Age-Related Macular Degeneration: A High Resolution Genome Scan for Susceptibility Loci in a Population Enriched for Late-Stage Disease.” It is available online at www.journals.uchicago.edu/AJHG/home.html.
In addition to Drs. Swaroop and Abecasis, other University of Michigan and Kellogg Eye Center study authors include Beverly M. Yashar, Yu Zhao, Noor M. Ghiasvand, Sepideh Zareparsi, Kari E. H. Branham, Adam C. Reddick, Edward H. Trager, Shigeo Yoshida, John Bahling, Elena Flippova, Susan G. Elner, Mark W. Johnson, Andrew K. Vine, and Julia E. Richards.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
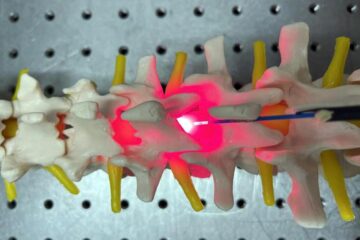
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
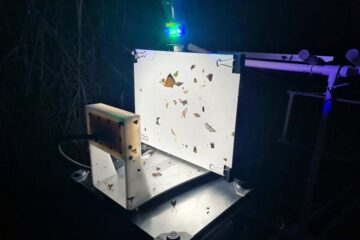

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….