Design of protein binders from target structure alone
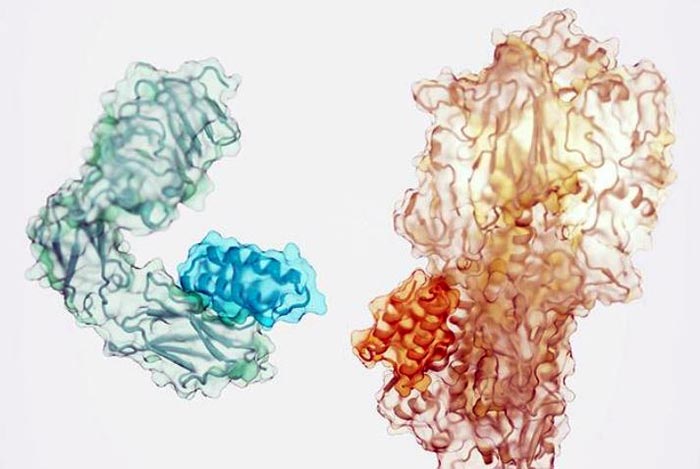
Small proteins (darker shade) designed to bind to the insulin receptor (left) and a component of the influenza virus (right).
Credit: Ian Haydon/Institute for Protein Design
A new method for generating potent, specific binding proteins yields candidate medicines for cancer, diabetes, inflammation and more.
A team of scientists has created a powerful new method for generating protein drugs. Using computers, they designed molecules that can target important proteins in the body, such as the insulin receptor, as well as vulnerable proteins on the surface of viruses. This solves a long-standing challenge in drug development and may lead to new treatments for cancer, diabetes, infection, inflammation, and beyond.
The research, appearing March 24 in the journal Nature, was led by scientists in the laboratory of David Baker, professor of biochemistry at the University of Washington School of Medicine and a recipient of the 2021 Breakthrough Prize in Life Sciences.
“The ability to generate new proteins that bind tightly and specifically to any molecular target that you want is a paradigm shift in drug development and molecular biology more broadly,” said Baker.
Antibodies are today’s most common protein-based drugs. They typically function by binding to a specific molecular target, which then becomes either activated or deactivated. Antibodies can treat a wide range of health disorders, including COVID-19 and cancer, but generating new ones is challenging. Antibodies can also be costly to manufacture.
A team led by two postdoctoral scholars in the Baker lab, Longxing Cao and Brian Coventry, combined recent advances in the field of computational protein design to arrive at a strategy for creating new proteins that bind molecular targets in a manner similar to antibodies. They developed software that can scan a target molecule, identify potential binding sites, generate proteins targeting those sites, and then screen from millions of candidate binding proteins to identify those most likely to function.
The team used the new software to generate high-affinity binding proteins against 12 distinct molecular targets. These targets include important cellular receptors such as TrkA, EGFR, Tie2, and the insulin receptor, as well proteins on the surface of the influenza virus and SARS-CoV-2 (the virus that causes COVID-19).
“When it comes to creating new drugs, there are easy targets and there are hard targets,” said Cao, who is now an assistant professor at Westlake University. “In this paper, we show that even very hard targets are amenable to this approach. We were able to make binding proteins to some targets that had no known binding partners or antibodies,”
In total, the team produced over half a million candidate binding proteins for the 12 selected molecular targets. Data collected on this large pool of candidate binding proteins was used to improve the overall method.
“We look forward to seeing how these molecules might be used in a clinical context, and more importantly how this new method of designing protein drugs might lead to even more promising compounds in the future,” said Coventry.
The research team included scientists from the University of Washington School of Medicine, Yale University School of Medicine, Stanford University School of Medicine, Ghent University, The Scripps Research Institute, and the National Cancer Institute, among other institutions.
This work was supported in part by The Audacious Project at the Institute for Protein Design, Open Philanthropy Project, National Institutes of Health (HHSN272201700059C, R01AI140245, R01AI150855, R01AG063845), Defense Advanced Research Project Agency (HR0011835403 contract FA8750-17-C-0219), Defense Threat Reduction Agency (HDTRA1-16-C-0029), Schmidt Futures, Gates Ventures, Donald and Jo Anne Petersen Endowment, and an Azure computing gift for COVID-19 research provided by Microsoft.
– Written by Ian Haydon, UW Medicine Institute for Protein Design
Journal: Nature
DOI: 10.1038/s41586-022-04654-9
Method of Research: Experimental study
Subject of Research: Not applicable
Article Title: Design of protein binding proteins from target structure alone
Article Publication Date: 24-Mar-2022
COI Statement: Competing interests Longxing Cao, Brian Coventry, Inns Gorenshink, , Buwei Hang, Nathaniel Bennet, Eva Maria Strauch., Lance Stewart and David Baker are co-inventors on a provisional patent application (21-0753-US-PRO) that incorporates discoveries described in this research news release and Nature paper.
Media Contact
Leila Gray
University of Washington School of Medicine/UW Medicine
leilag@u.washington.edu
Cell: 206-475-9809
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Webb captures top of iconic horsehead nebula in unprecedented detail
NASA’s James Webb Space Telescope has captured the sharpest infrared images to date of a zoomed-in portion of one of the most distinctive objects in our skies, the Horsehead Nebula….
Cost-effective, high-capacity, and cyclable lithium-ion battery cathodes
Charge-recharge cycling of lithium-superrich iron oxide, a cost-effective and high-capacity cathode for new-generation lithium-ion batteries, can be greatly improved by doping with readily available mineral elements. The energy capacity and…
Novel genetic plant regeneration approach
…without the application of phytohormones. Researchers develop a novel plant regeneration approach by modulating the expression of genes that control plant cell differentiation. For ages now, plants have been the…





















