Highly efficient, low-cost method developed to reduce DNA errors
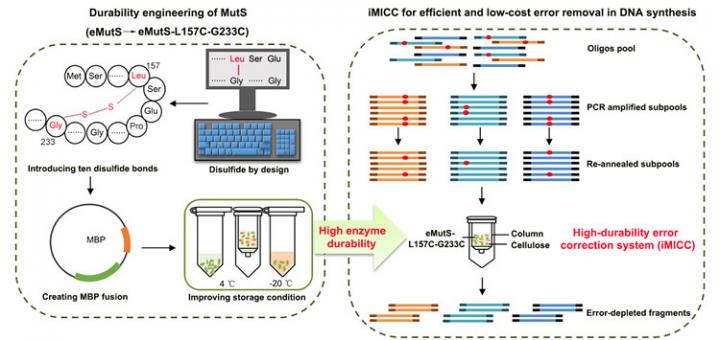
A newly developed low-cost method provides a highly efficient way to reduce DNA errors Credit: ZHANG Jia
Moreover, large-scale synthesis of DNA via such replication is a cornerstone of synthetic biology. However, just like the way we write or type an article, this DNA “writing” process is error prone, which has become a serious problem in large-scale DNA synthesis.
Now, a team of researchers from the Chinese Academy of Science (CAS) has developed a more efficient and cost-effective way to accurately synthesize DNA than traditionally used methods. They published their results on March 5, 2020 in ACS Synthetic Biology.
“In synthetic biology, genes, gene networks and even entire genomes are synthesized to create new functions,” said ZHANG Jia, paper first-author and a researcher from Single-Cell Center, Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT), CAS.
The synthetization process involves assembling DNA in liquid or on a microchip outfitted to intentionally pair specific genetic fragments. The problem, according to ZHANG, is that the fragments frequently mismatch and produce significant errors.
Current methods to reduce these errors involve a protein called MutS, which is attracted to mismatched genetic fragments. The protein acts as a flag on the errors, allowing scientists to identify and remove them.
This process is costly and time-consuming, however. One major reason is that, the key error-correcting enzyme of eMutS, a protein derived from E. coli and binds errors with high accuracy, is fragile and does not last long.
“To tackle this challenge, we have developed a simple, effective and cost-efficient error-correction system that is readily applicable in gene synthesis workflow,” ZHANG said.
The CAS team began by treating eMutS with chemical stabilizers in the form of a kind of molecular glue called disulfide bonds. With a strong chemical structure, the introduced bonds, plus improvements in enzyme production and storage, extended the life of the proteins from seven to 63 days.
Preparing the proteins for the error-removal process can take a significant amount of time, so this remarkable increase in enzyme durability means that researchers can go from preparing proteins once a week to once every two months. In industry-scale DNA synthesis workflows, this means significant reduction in operation, labor and time costs.
Furthermore, using the newly durable eMutS protein, 86.4% of the synthetic DNA fragments are completely free of errors – a nearly seven-fold increase in accuracy from the commercial enzyme systems currently on the market, according to ZHANG.
“This system's high fidelity, simple operation and low cost in error correction address one of the key challenges in DNA synthesis and could have implications for broad applications in synthetic biology, including industrial applications,” said XU Jian, senior author of the study and Director of Single-Cell Center, QIBEBT, CAS.
The team plans to continue improving the shelf-life of the proteins while also further increasing the accuracy of error removal in DNA synthesis.
###
This work was supported by the National Natural Science Foundation of China and National Science Fund for Distinguished Young Scholars.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
Microscopic basis of a new form of quantum magnetism
Not all magnets are the same. When we think of magnetism, we often think of magnets that stick to a refrigerator’s door. For these types of magnets, the electronic interactions…

An epigenome editing toolkit to dissect the mechanisms of gene regulation
A study from the Hackett group at EMBL Rome led to the development of a powerful epigenetic editing technology, which unlocks the ability to precisely program chromatin modifications. Understanding how…

NASA selects UF mission to better track the Earth’s water and ice
NASA has selected a team of University of Florida aerospace engineers to pursue a groundbreaking $12 million mission aimed at improving the way we track changes in Earth’s structures, such…