Stability for the genome
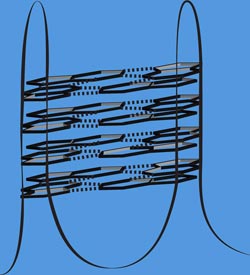
G-quadruplexes are node-like locations in DNA, where many guanines (nucleic acid) exercise a core function. They play an important role in the stability of the genome. Graphic: Katrin Paeschke<br>
In healthy human cells, the genome tends to be in a highly ordered state: DNA is neatly packed into precisely 46 chromosomes. In cancer cells, the picture is often very different: chromosomes may be broken or damaged in some other way, and sometimes even the entire genome is completely jumbled.
The stability of the genome depends on enzymes that have hardly changed over the course of evolution. This is what a biochemistry research group from the University of Würzburg, with colleagues from the USA and Switzerland, is reporting in the scientific journal “Nature”. The article also describes the DNA locations where enzymes exercise their effect.
Without helicases the genome breaks down
The enzymes concerned are so-called Pif1 helicases. “Whether in bacteria, yeast cells, or humans, Pif1 helicases or their homologs have the same function everywhere: they stabilize the genome,” says Dr. Katrin Paeschke, who runs an independent junior research group at the University of Würzburg’s Department of Biochemistry.
How important the helicases are becomes particularly apparent when they stop working. “This is often the case in breast cancer cells,” says Paeschke. There are also obvious consequences when the enzymes mutate in baker’s yeast cells: this leads to dramatic decomposition processes in their genome.
Where helicases stabilize DNA
Helicases exercise their stabilizing effect on special structures in the genome, the so-called G-quadruplexes. “These are node-like elements that can occur in the DNA molecule,” explains the Würzburg researcher. These structures require special protection: where they occur, DNA breaks down very easily. Without the protective helicases, particularly chaotic changes take place around the nodes, as Paeschke’s team observed.
Further research into G-quadruplexes
These new insights will not have an immediate impact on cancer treatment. “Looking further into the future, it is conceivable that regulating the G-quadruplexes might slow the breakdown of the genome in cancer cells,” comments Paeschke.
But first, the biochemist and her team are keen to investigate a different aspect: “G-quadruplexes are also misregulated in healthy cells in the presence of Pif helicases. We want to analyze how these important structures are repaired under these circumstances.”
About Katrin Paeschke
After a period of research at Princeton University (USA), Katrin Paeschke arrived at the University of Würzburg in early 2012. Since then she has been running an Emmy Noether junior research group here that is funded by the German Research Foundation (DFG). In 2012, she received a Heinz Maier-Leibnitz Prize from the DFG and the Röntgen Prize from the University of Würzburg. Both awards are bestowed upon outstanding young scientists.
“Pif1 family helicases suppress genome instability at G-quadruplex motifs”, Katrin Paeschke, Matthew L. Bochman, P. Daniela Garcia, Petr Cejka, Katherine L. Friedman, Stephen C. Kowalczykowski, and Virginia A. Zakian, Nature, 2013, May 8, doi: 10.1038/nature12149
Contact
Dr. Katrin Paeschke, Department of Biochemistry, Biocenter at the University of Würzburg, katrin.paeschke@biozentrum.uni-wuerzburg.de
Media Contact
More Information:
http://www.uni-wuerzburg.deAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

New insights into the mechanisms of chromosome segregation errors
Research on centromere structure… Researchers from the Kops group in collaboration with researchers from the University of Edinburgh, made a surprising new discovery in the structure of the centromere, a structure…

“Topological hall effect” in two-dimensional quantum magnets
In a recent study published in Nature Physics, researchers from the Hefei Institutes of Physical Science of the Chinese Academy of Sciences, together with researchers of University of Science and Technology of China, have introduced the…

Coating Technologies of the Future
LZH and Cutting Edge Coatings at Optatec… At Optatec 2024, the Laser Zentrum Hannover e.V. (LZH) and Cutting Edge Coatings GmbH (CEC) present new opportunities in coating technologies for the…