Scientists watch cell-shape process for first time

All plant and animal cells rely on an elaborate array of molecular rods built from the protein tubulin. These rods, called microtubules, organize the cell and generate forces needed to support cell shape, cell movement, and importantly, cell division.
To perform these tasks, microtubules need to be organized into specific configurations. Animal cells separate their chromosomes during cell division by organizing the microtubules network from centrioles. A big mystery is how plants, which do not have centrioles organize their microtubule network. Understanding these mechanisms of molecular organization is a primary goal of cell biology.
As co-author David Ehrhardt from Carnegie's Department of Plant Biology explained: “In many cells, microtubule arrays are created with aid of a centralized body called a centrosome. Centrosomal arrays have been a focus of research for decades and much is now understood about how these arrays are created and organized by the centrosome. However, many differentiated animal cells, and flowering plant cells have arrays that are created independently of a centrosome. In fact, flowering plants lack centrosomes all together. Although these centrosome arrays are common in nature, they have received less study and their organization mechanisms remain largely mysterious.”
The Ehrhardt lab previously found that individual microtubules in plant cell arrays are born at many locations along the inside of the cell membrane, where they are detached from the sites of birth and move along the membrane to interact with other microtubules. A primary challenge for investigating the molecular basis for these processes has been visualization of the protein complexes that give birth to new microtubule polymers.
The Ehrhardt and Hashimoto groups met this challenge by tagging a component of these complexes, known as nucleating complexes, with multiple copies of a fluorescent protein derived from jellyfish. When introduced into plant cells and visualized with highly sensitive spinning disk confocal microscopy, this tagged protein permitted the researchers to observe what happens as the microtubule array is being built.
Ehrhardt continued: “In centrosomal arrays, these nucleating complexes are recruited to the centrosome, where they give rise to a star-shaped array centered near the nucleus. By contrast, in the cells we studied these complexes were distributed at the cell membrane and were primarily located along the sides of other microtubules, an association that was correlated with their activity. So, microtubules appear to be important for locating and regulating their own formation proteins. In addition, daughter microtubules were created either at a distinct angle to the mother polymer, or in parallel to it. This choice of angle may play a role in either creating new organizational states or maintaining an existing one.”
The investigators observed that formation complexes frequently did not remain in place after creating new microtubules, but often left, presumably to go through a new cycle of microtubule creation at a new location. The scientists hypothesized that liberation of the complexes from mother microtubules might be related to the mechanism of daughter microtubule detachment from origination sites.
To explore these questions, the investigators introduced their probe into a mutant lacking the protein katanin (named for a Japanese word for sword), whose job it is to cut microtubules into pieces. The scientists thought that katanin might be responsible for separating new microtubules from their formation complexes. In fact, without the cutting protein, the daughter microtubules completely failed to detach from their birth sites, and tagged formation complexes remained at the base of the daughter microtubule. The only time they saw a formation complex leave in the mutants was when the microtubule completely depolymerized—that is, the process whereby a large molecule decomposes into individual units. When this occurred, the tagged complex also disappeared. The results indicate that the formation complexes remain associated with mother microtubules until the daughter microtubule is removed either by katanin cutting or by complete depolymerization.
“As far as we are aware, this research is the first to witness the dynamics of individual gamma tubulin complex processes, which are fundamental to every plant and animal,” remarked Ehrhardt. “We look at our plant system as a model for non-centrosomal array organization, which also occurs in many important differentiated animal cells. While we anticipate that some of the molecular players may be different, many of the principles may be similar. What we learn here could help us understand basic mechanisms underlying crop plant growth and development, and could have implications for understanding the process of acquiring cell shape and function of human cells.”
*Authors on the paper are Masayoshi Nakamura of the Nara Institute of Science and Technology; David Ehrhardt of Carnegie; and Takashi Hashimoto of Nara and Stanford University. The work was partially supported by the Carnegie Institution for Science, the Japanese NARA Institute of Science and Technology and the Ministry of Education, Culture, Sports, Science and Technology.
The Carnegie Institution for Science (carnegiescience.edu) is a private, nonprofit organization headquartered in Washington, D.C., with six research departments throughout the U.S. Since its founding in 1902, the Carnegie Institution has been a pioneering force in basic scientific research. Carnegie scientists are leaders in plant biology, developmental biology, astronomy, materials science, global ecology, and Earth and planetary science.
Media Contact
More Information:
http://www.carnegiescience.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
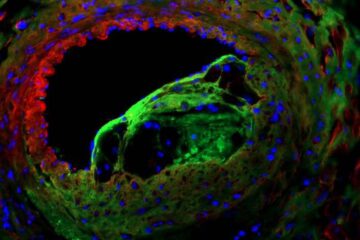
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…





















