Scientists solve 40-year mystery of how sodium controls opioid brain signaling
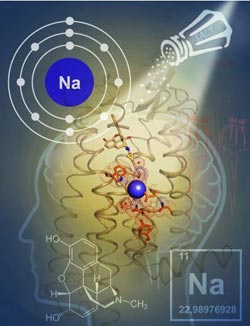
Researchers have unraveled the mystery of how the element sodium influences the signaling of opioid receptors, opening the door to new kinds of therapies.<br><br>Credit: Photo courtesy of the Scripps Research Institute<br>
Scientists have discovered how the element sodium influences the signaling of a major class of brain cell receptors, known as opioid receptors. The discovery, from The Scripps Research Institute (TSRI) and the University of North Carolina (UNC), suggests new therapeutic approaches to a host of brain-related medical conditions.
“It opens the door to understanding opioid related drugs for treating pain and mood disorders, among others,” said lead author Dr. Gustavo Fenalti, a postdoctoral fellow in the laboratory of Professor Raymond C. Stevens of TSRI's Department of Integrative Structural and Computational Biology.
“This discovery has helped us decipher a 40-year-old mystery about sodium's control of opioid receptors,” said Stevens, who was senior author of the paper with UNC pharmacologist Professor Bryan Roth. “It is amazing how sodium sits right in the middle of the receptor as a co-factor or allosteric modulator.”
The findings appear in an advanced online publication in the journal Nature on January 12, 2014.
A Sharper Image
The researchers revealed the basis for sodium's effect on signaling with a high-resolution 3-D view of an opioid receptor's atomic structure. Opioid receptors are activated by peptide neurotransmitters (endorphins, dynorphins and enkephalins) in the brain. They can also be activated by plant-derived and synthetic drugs that mimic these peptides: among them morphine, codeine, oxycodone and heroin.
Despite these receptors' crucial importance in health and disease, including pain disorders and addictions, scientists have only begun to understand in detail how they work. Opioid receptors are inherently flimsy and fragile when produced in isolation, and thus have been hard to study using X-ray crystallography, the usual structure-mapping method for large proteins.
In recent years, the Stevens laboratory has helped pioneer the structure determination of G protein-coupled receptors. Although the first crystallographic structures of opioid receptors were determined in 2012, these structural models weren't fine-grained enough to solve a lingering mystery, particularly for the human delta opioid receptor.
That mystery concerned the role of sodium. The element is perhaps best known to biologists as one of the key “electrolytes” needed for the basic workings of cells. In the early 1970s, researchers in the laboratory of neuroscientist Solomon Snyder at Johns Hopkins University, who had helped discover opioid receptors, found evidence that sodium ions also act as a kind of switch on opioid receptor signaling. They noted that at concentrations normally found in brain fluid, these ions reduced the ability of opioid peptides and drugs like morphine to interact with opioid receptors.
How sodium could exert this indirect (“allosteric”) effect on opioid receptor activity was unclear—and has remained an unsolved puzzle for decades. Now that scientists have discovered the mechanism of sodium's effect, then in principle they can exploit it to develop better opioid-receptor-targeting drugs.
A Switch Controlling Pain, Depression and Mood Disorders
For the new study, the team constructed a novel, fusion-protein-stabilized version of one of the main opioid receptors in the human brain, known as the delta opioid receptor, and managed to form crystals of it for X-ray crystallography. The latter revealed the receptor's 3-D atomic structure to a resolution of 1.8 Angstroms (180 trillionths of a meter)—the sharpest picture yet of an opioid receptor.
“Such a high resolution is really necessary to be able to understand in detail how the receptor works,” said Stevens.
The analysis yielded several key details of opioid receptor structure and function, most importantly the details of the “allosteric sodium site,” where a sodium ion can slip in and modulate receptor activity.
The team was able to identify the crucial amino acids that hold the sodium ion in place and transmit its signal-modulating effect. “We found that the presence of the sodium ion holds the receptor protein in a shape that gives it a different affinity for its corresponding neurotransmitter peptides,” Fenalti said.
With the structural data in hand, the researchers designed new versions of the receptor, in which key sodium-site amino-acids were mutated, to see how this would affect receptor signaling. Co-lead author Research Associate Patrick M. Giguere and colleagues in Roth's Laboratory at UNC, which has long collaborated with the Stevens laboratory, tested these mutant receptors and found that certain amino-acid changes cause radical shifts in the receptor's normal signaling response.
The most interesting shifts involved a little-understood secondary or “alternative” signaling route, known as the beta-arrestin pathway, whose activity can have different effects depending on the type of brain cell involved. Some drugs that normally bind to the delta opioid receptor and have little or no effect on the beta-arrestin pathway turned out to strongly activate this pathway in a few of these mutant receptors.
In practical terms, these findings suggests a number of ways in which new drugs could target these receptors—and not only delta opioid receptors but also the other two “classical” opioid receptors, mu and kappa opioid receptors. “The sodium site architecture and the way it works seems essentially the same for all three of these opioid receptor types,” said Fenalti.
Other co-authors of the study, “Molecular control of Delta-opioid receptor signaling,” were Assistant Professor Vsevolod Katritch, former Staff Scientist Aaron A. Thompson and Associate Professor Vadim Cherezov of TSRI, and Assistant Scientific Director Xi-Ping Huang of the University of North Carolina.
The research was funded in part by the National Institutes of Health (grants P50 GM073197, U54 GM094618, R01 DA017204), the National Cancer Institute (Y1-CO-1020), the National Institute for General Medical Sciences (Y1-GM-1104) and the National Institute of Mental Health Psychoactive Drug Screening Program.
Media Contact
More Information:
http://www.scripps.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
Microscopic basis of a new form of quantum magnetism
Not all magnets are the same. When we think of magnetism, we often think of magnets that stick to a refrigerator’s door. For these types of magnets, the electronic interactions…

An epigenome editing toolkit to dissect the mechanisms of gene regulation
A study from the Hackett group at EMBL Rome led to the development of a powerful epigenetic editing technology, which unlocks the ability to precisely program chromatin modifications. Understanding how…

NASA selects UF mission to better track the Earth’s water and ice
NASA has selected a team of University of Florida aerospace engineers to pursue a groundbreaking $12 million mission aimed at improving the way we track changes in Earth’s structures, such…