Beyond the DNA: Chemical signatures reveal genetic switches in the genome

Investigators from the Ludwig Institute for Cancer Research (LICR) and the University of California, San Diego (UCSD) School of Medicine have made a breakthrough in identifying functional elements in the human genome, according to a report published online today in Nature Genetics.
While the DNA sequence can identify genes (the 'what') within the genome, it cannot answer the more fundamental questions of 'how,' 'when' and 'where' gene products are expressed. However, the LICR team and collaborators have developed a novel method to identify and predict the 'promoter' and 'enhancer' regions that switch on transcription, the first step in gene expression. This study is an important step towards large-scale functional annotation of 'enhancers,' which establish the rate at which gene expression occurs and determine the tissues in which genes are expressed.
“The human genome is packaged within each cell by chromatin, a structure composed of DNA wrapped around proteins called histones,” explains lead author Nathaniel Heintzman, from LICR and the UCSD Biomedical Sciences Graduate Program. “We used genome-scale approaches to analyze the chromatin architecture in human cells and discovered distinct signatures of modified histones that correspond to known promoters and enhancers. These findings enabled us to generate computational algorithms that identified hundreds of novel genomic regions with regulatory potential.” Moreover, says Heintzman, this 'histone code' was able to accurately distinguish between promoters, from which gene transcription is directly initiated, and enhancer regions – some of which are very distant from the target gene – to which transcription-activating cofactors bind.
According to LICR's Dr. Bing Ren, the study's senior author and assistant professor at UCSD's Department of Cellular and Molecular Medicine, the method is widely applicable and its unbiased approach is likely to yield new information on gene expression and how it is altered in disease. “The beauty of this approach is that it utilizes a chemical signature present on histones but not DNA. Existing methods to predict enhancers rely on the DNA sequence alone, but these are inadequate because we have an incomplete understanding of the sequence features that identify enhancers. The elucidation of a common histone modification signature will enable scientists to quickly identify enhancers and promoters for a gene, which will lead in turn to rapid identification of factors that control its expression.” This method might also be used to identify the disruptions in gene networks that occur during cancer progression, says Dr. Ren, which might lead to novel strategies for cancer diagnosis.
Media Contact
More Information:
http://www.licr.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
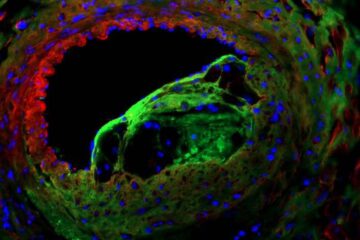
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…





















