Breakthrough in detecting DNA mutations could help treat tuberculosis, cancer

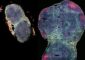
This conceptual image shows probe and target complexes at different stages of the reaction that checks for mutations. The red dots represent mutations in a target base pair, while sequences with illuminated green lights indicate that no mutation was found in the reaction.<br><br>Credit: Yan Liang, L2XY2.com<br>
Now, researchers have developed a new method that can look at a specific segment of DNA and pinpoint a single mutation, which could help diagnose and treat diseases such as cancer and tuberculosis.
These small changes can be the root of a disease or the reason some infectious diseases resist certain antibiotics. The findings were published online this week (July 28) in the journal Nature Chemistry.
“We've really improved on previous approaches because our solution doesn't require any complicated reactions or added enzymes, it just uses DNA,” said lead author Georg Seelig, a University of Washington assistant professor of electrical engineering and of computer science and engineering. “This means that the method is robust to changes in temperature and other environmental variables, making it well-suited for diagnostic applications in low-resource settings.”
DNA is a type of nucleic acid, the biological molecule that gives all living things their unique genetic signatures. In a double strand of DNA, known as a double helix, a series of base pairs bond and encode our genetic information. As genomics research has progressed, it's clear that a change of just one base pair – a sequence mutation, an insertion or a deletion – is enough to trigger major biological consequences. This could explain the onset of disease, or the reason some diseases don't respond to usual antibiotic treatment.
Take, for example, tuberculosis – a disease that's known to have drug-resistant strains. Its resistance to antibiotics often is due to a small number of mutations in a specific gene. If a person with tuberculosis isn't responding to treatment, it's likely because there is a mutation, Seelig said.
Now, researchers have the ability to check for that mutation preventatively.
Seelig, along with David Zhang of Rice University and Sherry Chen, a UW doctoral student in electrical engineering, designed probes that can pick out mutations in a single base pair in a target stretch of DNA. The probes allow researchers to look in much more detail for variations in long sequences – up to 200 base pairs – while current methods can detect mutations in stretches of up to only 20.
“In terms of specificity, our research suggests that we can do quadratically better, meaning that whatever the best level of specificity, our best will be that number squared,” said Zhang, an assistant professor of bioengineering at Rice University.
The testing probes are designed to bind with a sequence of DNA that is suspected of having a mutation. The researchers do this by creating a complimentary sequence of DNA to the double-helix strand in question. Then, they allow molecules containing both sequences to mix in a test tube in salt water, where they naturally will match up to one another if the base pairs are intact. Unlike previous technologies, the probe molecule checks both strands of the target double helix for mutations rather than just one, which explains the increased specificity.
The probe is engineered to emit a fluorescent glow if there's a perfect match between it and the target. If it doesn't illuminate, that means the strands didn't match and there was in fact a mutation in the target strand of DNA.
The researchers have filed a patent on the technology and are working with the UW Center for Commercialization. They hope to integrate it into a paper-based diagnostic test for diseases that could be used in parts of the world with few medical resources.
The research was funded by the National Institutes of Health, the National Science Foundation and the Department of Defense's Advanced Research Projects Agency.
For more information, contact Seelig at gseelig@uw.edu.
Media Contact
More Information:
http://www.uw.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
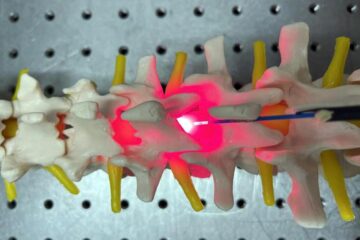
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
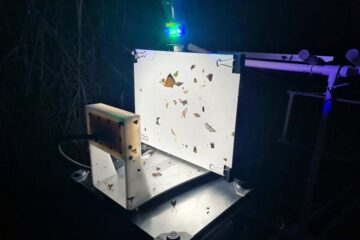

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….