Mustard-root map breaks new ground tracking gene expression

New ’global’ technique a dividend of NSF’s Arabidopsis 2010 effort
A new “gene expression” map is helping scientists track how a complex tissue ultimately arises from the blueprint of thousands of genes.
Focusing on the root of a small flowering mustard plant, Arabidopsis thaliana, a research team led by Duke University biologist Philip Benfey created a detailed mosaic of cells showing where and when about 22,000 of the plant’s roughly 28,000 genes are activated within growing root tissue.
The results, announced in the Dec. 12th issue of the journal Science, are the first to demonstrate “this level of resolution of gene expression on a global basis for any organism,” said Benfey. The work, he said, serves as “a proof of principle” that similar approaches can be applied to other plant organs and other organisms.
It also marks the first time researchers have tracked the vast majority of an organism’s genes as they are switched on and off as cells grow, continually divide and ultimately differentiate to build specialized tissue.
The ability to track gene expression on this scale (with each cellular division along a comprehensive front) is critical to answering one of biology’s basic, yet most puzzling, processes: How do distinct, yet coordinated organs and specialized cells arise from the endless division of cells that initially seemed quite similar? For example, how does this complex process with a simple name, development, begin with a single, fertilized cell and ultimately yield a plant with roots, leaves, buds and blooms?
The researchers also found that different types of root cells tended to express particular sets of genes that were clustered together on the plant chromosomes. Understanding these patterns of cell types and gene clusters, Benfey said, could help biologists decipher the genetic machinery of development and eventually lead to new ways to enhance crops.
The research was funded by the National Science Foundation (NSF), the independent federal agency that supports fundamental research and education across all fields of science and engineering.
Three years ago following an international effort, Arabidopsis became the first plant to have its genome sequence completed. NSF, a key funder of the sequencing effort, then launched “Arabidopsis 2010,” a program to determine the function of the all of the plant’s genes in this decade. (It, too, is part of a multinational effort.)
The gene-expression map announced in Science resulted from a $2.2 million 2010 project to apply “genomics approaches to finding transcriptional networks.”
(Using a gene’s DNA as the template, the transcription process creates strands of RNA, molecules that control the building of proteins and serve as catalysts. A network of various biochemical factors, such as signaling hormones, can affect this process.)
According to Joanne Tornow, a program director in NSF’s Division of Molecular and Cellular Biosciences, “the creation of the root map is a terrific advance forward.”
“The process should work with other plant tissues, although beyond the root it may be more difficult to observe changes in gene expression over developmental time,” said Tornow.
“But this lays the groundwork for looking at how various biological pathways interlink in transcriptional networks,” she said. “There are still thousands of genes in Arabidopsis, and we know almost nothing about their function. By knowing when a gene is expressed and where it is expressed, we get clues about the processes it is involved with and potentially its function as well.”
To develop the map, Benfey worked with colleagues at Duke, New York University and the University of Arizona. In Science, they report, “High throughput techniques allowed the harvesting, protoplasting (breaking down of cell walls by enzymes), and sorting of approximately 10 million cells in about 1.5 hours.”
To track gene expression over time, they relied upon the fact that a root cell’s advancing stages of development correlate to its distance from the root tip’s growing point.
To track the lineage of individual cells as they developed into specific tissue, they attached marker genes to genes characteristic of each of five different cell types or tissues. The marker genes produce a tell-tale, and therefore traceable, green fluorescent protein (GFP) when the gene they’re attached to is activated.
Then, using methods invented by David Galbraith at the University of Arizona, researchers moved quickly to sort, isolate and identify the fluorescence-activated genes, which glow under ultraviolet light when the gene they’ve marked is being expressed. They conducted the process during three successive stages synchronously across five zones of cells and tissues in the root.
To generate a visual map of 15 “subgrids,” the massive amount of data was “digitally reconstructed” with the intensity of gene expression illustrated along a color scale.
According to Benfey, “other genomic studies, in which whole tissues were ground up and their global gene expression profiles determined, certainly generated much useful information. However, critical information on the mechanisms of development was lost. Development occurs at the single cell level, and there’s a dramatic difference from one cell to the next, in terms of its gene expression.”
NSF Program Officer: Joanne Tornow, Genes and Genome Systems, 703-292-8439, jtornow@nsf.gov.
Principal Investigator: Philip Benfey, Department of Biology, Duke University, 919-613-8182, philip.benfey@duke.edu.
Additional Investigator: David Galbraith, Professor of Plant Sciences, University of Arizona, 520-621-9153, galbraith@arizona.edu.
Related news releases from Duke University:
http://www.dukenews.duke.edu/news/newsrelease.asp?id=3053&catid=2,46&cpg=newsrelease.asp. Contact at Duke: Dennis Meredith, 919-681-8054, dennis.meredith@duke.edu.
Media Contact
More Information:
http://www.nsf.gov/od/lpa/news/03/pr03139.htmAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Why getting in touch with our ‘gerbil brain’ could help machines listen better
Macquarie University researchers have debunked a 75-year-old theory about how humans determine where sounds are coming from, and it could unlock the secret to creating a next generation of more…

Attosecond core-level spectroscopy reveals real-time molecular dynamics
Chemical reactions are complex mechanisms. Many different dynamical processes are involved, affecting both the electrons and the nucleus of the present atoms. Very often the strongly coupled electron and nuclear…
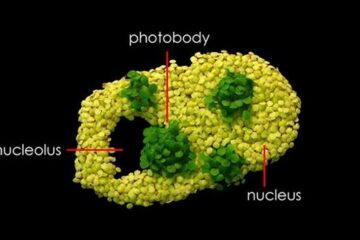
Free-forming organelles help plants adapt to climate change
Scientists uncover how plants “see” shades of light, temperature. Plants’ ability to sense light and temperature, and their ability to adapt to climate change, hinges on free-forming structures in their…