Amping antimicrobial discovery with automation
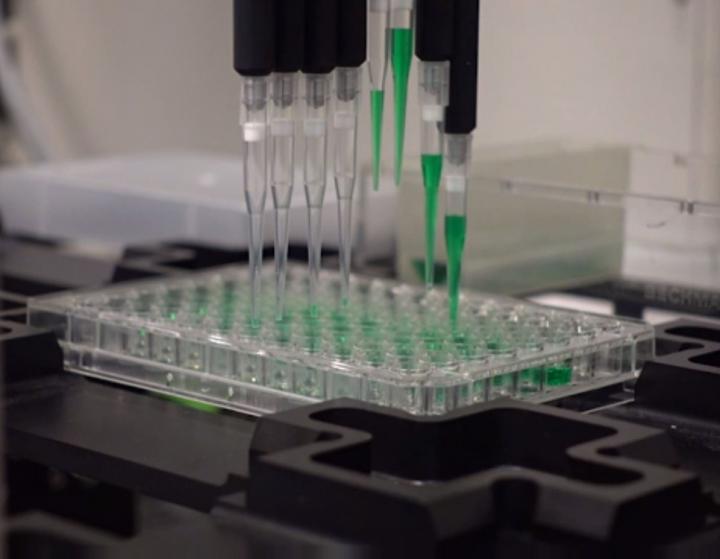
A robotic device designed for high-throughput screening of drug toxicity also could also accelerate antimicrobial discovery, NIST researchers report. Credit: Stoughton/NIST
The antimicrobial arsenal that we count on to save millions of lives each year is alarmingly thin–and these microbes are rapidly evolving resistance to our weapons. But help may be on the way: In a study posted in the AMB Express, researchers from the National Institute of Standards and Technology (NIST) show that automated techniques commonly used to screen new drugs for mammalian cell toxicity could also dramatically speed up the challenging task of antimicrobial discovery.
In the age-old struggle between humans and microbes, bacteria seem to be regaining the offensive. Only around a dozen classes of chemicals protect us from the myriad pathogens that populate our environment. Numerous agencies, including the World Health Organization and the Centers for Disease Control and Prevention, have recently warned that evolved resistance could soon render common antibiotics useless, and that few replacement drugs are in the pipeline.
The shortage of new antimicrobials is not a result of scientists lacking candidate chemicals. The fungal and plant worlds abound with potential antimicrobials, and chemists concoct new synthetic molecules all the time. However, a major bottleneck occurs at the lab bench. Any candidate compound must be tested at multiple concentrations against multiple strains of bacteria in different forms. This remains a cumbersome process, with numerous time- and labor-intensive steps that lab workers must currently carry out by hand.
But NIST researcher Samuel Forry and colleagues are convinced that the process could be vastly sped up using automation. To do so, Forry and his team looked to one of the pharmaceutical industry's most powerful tools: high-throughput screening. For several decades, companies have routinely used automated systems to test potential drugs' effects on mammalian cells in culture. In these studies, robots prepare samples of cells in arrays of small plastic wells, inject measured amounts of drugs and test whether cells live or die. The method can quickly assess multiple chemicals at different concentrations, all in parallel and with minimal human intervention.
High-throughput screening has seen limited use for antimicrobial discovery, Forry says, because less research and development money is available and because of the large variation among microbial populations and growth conditions. Hoping to stimulate the field, Forry and his team adapted a high-throughput screening robot for antimicrobial testing. The researchers tested a set of antimicrobial compounds known as pyridinium salts against the common bacterium Streptococcus mutans, which causes tooth decay.
Part of the challenge in identifying useful antimicrobial compounds is that chemicals that kill free-swimming cells are often less effective against the same bacteria growing in biofilms like the plaque that can form on teeth. So Forry's team used automation to culture both free-swimming cells and biofilms, as well as an intermediate state, side-by-side in 96-well plates. The researchers measured antimicrobial activity in three different ways by identifying the concentrations that reduced bacterial activity by half, that prevented any detectable activity, and that entirely killed the bacteria. They determined the drugs' effects with high throughput by measuring light passing through the wells or using chemicals that change color to indicate metabolic activity.
The team found that the automated system delivered results indistinguishable from those obtained by doing the experiments by hand. More importantly, the robot took only a third as much time as humans do, freed up laboratory personnel for other tasks, and carried out the procedures without errors. “That's a huge improvement from the point of view of laboratory workflow and a great boon for people trying to identify and characterize antimicrobials,” Forry says.
The trials weren't fully automated–for instance, the researchers moved samples from the incubator to the screening robot by hand–but Forry says his team has demonstrated the concept, and existing technology can fill in the remaining steps. He expects other research labs will adopt the technology first, followed by pharmaceutical companies. “Once a number of people start to use this and find that it works for them as well as it has worked for us, I could easily see companies and contract labs doing it.”
###
Paper: S.P. Forry, M.C. Madonna, D. López-Pérez, N.J. Lin and M.D. Pasco, “Automation of Antimicrobial Activity Screening,” AMB Express 2016, 6:20. DOI: 10.1186/s13568-016-0191-2
Media Contact
All latest news from the category: Health and Medicine
This subject area encompasses research and studies in the field of human medicine.
Among the wide-ranging list of topics covered here are anesthesiology, anatomy, surgery, human genetics, hygiene and environmental medicine, internal medicine, neurology, pharmacology, physiology, urology and dental medicine.
Newest articles

Why getting in touch with our ‘gerbil brain’ could help machines listen better
Macquarie University researchers have debunked a 75-year-old theory about how humans determine where sounds are coming from, and it could unlock the secret to creating a next generation of more…

Attosecond core-level spectroscopy reveals real-time molecular dynamics
Chemical reactions are complex mechanisms. Many different dynamical processes are involved, affecting both the electrons and the nucleus of the present atoms. Very often the strongly coupled electron and nuclear…
Free-forming organelles help plants adapt to climate change
Scientists uncover how plants “see” shades of light, temperature. Plants’ ability to sense light and temperature, and their ability to adapt to climate change, hinges on free-forming structures in their…