Researchers get unprecedented view of protein folding that may help develop brain disease therapies

Misfold an origami swan and the worst that happens is you wind up with an ugly paper duckling. Misfold one of the vital proteins in your body – each of which must be folded in a particular way to perform its function – and the result can be a debilitating neurodegenerative disease such as Alzheimer's or Huntington's.
There are no cures for such brain-wasting diseases, but now Stanford researchers have taken an important step that may one day aid in developing therapies for them. They have literally popped the lid off one of the microscopic chambers in which many of life's most crucial proteins are folded, witnessing a surprising mechanism as the heretofore hidden folding process happened before their eyes.
Virtually all proteins need to be folded, whether in primitive organisms such as bacteria or multicellular creatures such as humans. Many are guided through the process by molecules called chaperones, of which a specialized subset – chaperonins – folds many of the most complex proteins.
Folding in bacteria has been studied in detail, but Judith Frydman, a professor of biology who led the Stanford research, said this is the first time anyone has seen the folding process performed in higher organisms.
“The mechanism of folding we saw in the chaperonin is very different from what we expected and from what has been seen in bacteria,” Frydman said. “It was really surprising, and we are still amazed that it worked. This chaperonin appears to provide a unique chemical environment.”
Chaperonins are shaped like a barrel, with two ring-shaped chambers arranged one atop the other. At the open end of each ring is a lid that opens and closes in a spiraling fashion, like the aperture of a camera, something Frydman's team discovered in 2008 while studying the chaperonin called TRiC. Since then, they've been working to solve the puzzle of how a protein gets folded once the chaperonin has grabbed it, pulled it into the chamber and the aperture has closed. A paper describing their findings was published earlier this year in Cell.
Frydman said there were two likely ways in which a protein, initially a linear chain of molecules (amino acids), could theoretically be folded inside the chamber.
One is by mechanical means, with the chamber holding onto the protein and physically pushing it into the right shape.
“The other one is that when the lid closes, the chaperonin lets go of the protein, but some special chemical properties in this chamber somehow make it fold,” she said. “Our evidence is that this mechanism is the correct one.”
The only way to know which mechanism was doing the work was to see inside the chamber while the folding was happening, but simply opening up the lid wouldn't work, because the shape of the entire chamber changes in accordance with the motion of the lid. When the lid spirals open, the walls of the chamber spiral open, too, and the protein floats away.
To see what was happening, Frydman's team devised a chemical “trick” by which they could remove the lid on the chamber, but still get the walls of the chamber to close in, as if the lid were spiraling.
When they “closed” the lidless chamber, the chaperonin simply released the protein that had been destined to be folded. Like a long balloon that slipped from a child's grip before it could be folded into a giraffe, the protein simply drifted off.
The challenge then became figuring out how the protein was getting released.
“One of the reasons why the mechanical model of pushing the protein into shape without letting go had been proposed was because there was no obvious way for this chaperonin to let go of the protein,” Frydman said.
When a protein gets grabbed for folding by TRiC, it is held by eight binding sites along the walls of the chamber. Between each binding site is a tiny loop. Frydman's team suspected that during the closing process, the loops might move to somehow “shave off” the protein and release it into the folding chamber. One of her students made mutations in the loop. When the researchers did experiments in which TRiC chaperonins equipped with mutated loops were closed, the protein stayed put. It also failed to fold.
“That suggests that the way this chaperonin folds its proteins is by releasing them in a closed chamber that has very special chemical properties,” Frydman said.
“This mechanism of release is completely different from what has been seen in any other chaperone. That was very, very surprising.”
The experimental work described in the Cell paper was done using a simpler version of TRiC, from a single-celled organism, than would be found in multi-cellular organisms, Frydman said, because the simpler version is much easier to manipulate.
“Now we are interested in going back to the eukaryotic [multi-cellular] complex, where every binding site in the folding chamber is different and every release loop is different,” Frydman said. “I think this really opens up a lot of interesting avenues to explore how this works in higher organisms. Since TRiC helps fold many disease-linked proteins, and is central to protect cells from misfolding diseases such as Huntington's disease, this work could have many therapeutic applications.”
Media Contact
More Information:
http://www.stanford.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
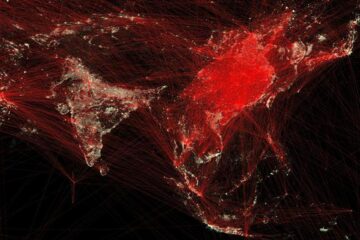
Economies take off with new airports
A global study by an SUTD researcher in collaboration with scientists from Japan explores the economic benefits of airport investment in emerging economies using nighttime satellite imagery. Be it for…
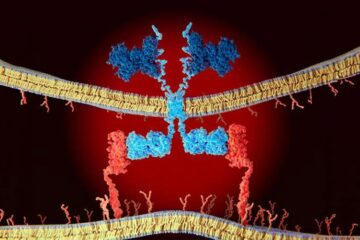
CAR T–cell immunotherapy targets
Pan-cancer analysis uncovers a new class of promising CAR T–cell immunotherapy targets. Scientists at St. Jude Children’s Research Hospital found 156 potential CAR targets across the brain and solid tumors,…

Stony coral tissue loss disease
… is shifting the ecological balance of Caribbean reefs. The outbreak of a deadly disease called stony coral tissue loss disease is destroying susceptible species of coral in the Caribbean…





















