One species' entire genome discovered inside another's

Scientists at the University of Rochester and the J. Craig Venter Institute have discovered a copy of the entire genome of a bacterial parasite residing inside the genome of its host species.
The finding, reported in today’s Science, suggests that lateral gene transfer—the movement of genes between unrelated species—may happen much more frequently between bacteria and multicellular organisms than scientists previously believed, posing dramatic implications for evolution.
Such large-scale heritable gene transfers may allow species to acquire new genes and functions extremely quickly, says Jack Werren, a principle investigator of the study.
The results also have serious repercussions for genome-sequencing projects. Bacterial DNA is routinely discarded when scientists are assembling invertebrate genomes, yet these genes may very well be part of the organism’s genome, and might even be responsible for functioning traits.
“This study establishes the widespread occurrence and high frequency of a process that we would have dismissed as science fiction until just a few years ago,” says W. Ford Doolittle, Canada Research Chair in Comparative Microbial Genomics at Dalhousie University, who is not connected to the study. “This is stunning evidence for increased frequency of gene transfer.”
“It didn’t seem possible at first,” says Werren, professor of biology at the University of Rochester and a world-leading authority on the parasite, called Wolbachia. “This parasite has implanted itself inside the cells of 70 percent of the world’s invertebrates, coevolving with them. And now, we’ve found at least one species where the parasite’s entire or nearly entire genome has been absorbed and integrated into the host’s. The host’s genes actually hold the coding information for a completely separate species.”
Wolbachia may be the most prolific parasite in the world—a “pandemic,” as Werren calls it. The bacterium invades a member of a species, most often an insect, and eventually makes its way into the host’s eggs or sperm. Once there, the Wolbachia is ensured passage to the next generation of its host, and any genetic exchanges between it and the host also are much more likely to be passed on.
Since Wolbachia typically live within the reproductive organs of their hosts, Werren reasoned that gene exchanges between the two would frequently pass on to subsequent generations. Based on this and an earlier discovery of a Wolbachia gene in a beetle by the Fukatsu team at the University of Tokyo, Japan, the researchers in Werren’s lab and collaborators at J. Craig Venter Institute (JCVI) decided to systematically screen invertebrates. Julie Dunning-Hotopp at JCVI found evidence that some of the Wolbachia genes seemed to be fused to the genes of the fruitfly, Drosophila ananassae, as if they were part of the same genome.
Michael Clark, a research associate at Rochester then brought a colony of ananassae into Werren’s lab to look into the mystery. To isolate the fly’s genome from the parasite’s, Clark fed the flies a simple antibiotic, killing the Wolbachia. To confirm the ananassae flies were indeed cured of the wolbachia, Clark tested a few samples of DNA for the presence of several Wolbachia genes.
To his dismay, he found them.
“For several months, I thought I was just failing,” says Clark. “I kept administering antibiotics, but every single Wolbachia gene I tested for was still there. I started thinking maybe the strain had grown antibiotic resistance. After months of this I finally went back and looked at the tissue again, and there was no Wolbachia there at all.”
Clark had cured the fly of the parasite, but a copy of the parasite’s genome was still present in the fly’s genome. Clark was able to see that Wolbachia genes were present on the second chromosome of the insect.
Clark confirmed that the Wolbachia genes are inherited like “normal” insect genes in the chromosomes, and Dunning-Hotopp showed that some of the genes are “transcribed” in uninfected flies, meaning that copies of the gene sequence are made in cells that could be used to make Wolbachia proteins.
Werren doesn’t believe that the Wolbachia “intentionally” insert their genes into the hosts. Rather, it is a consequence of cells routinely repairing their damaged DNA. As cells go about their regular business, they can accidentally absorb bits of DNA into their nuclei, often sewing those foreign genes into their own DNA. But integrating an entire genome was definitely an unexpected find.
Werren and Clark are now looking further into the huge insert found in the fruitfly, and whether it is providing a benefit. “The chance that a chunk of DNA of this magnitude is totally neutral, I think, is pretty small, so the implication is that it has imparted of some selective advantage to the host,” says Werren. “The question is, are these foreign genes providing new functions for the host” This is something we need to figure out.”
Evolutionary biologists will certainly take note of this discovery, but scientists conducting genome-sequencing projects around the world also may have to readjust their thinking.
Before this study, geneticists knew of examples where genes from a parasite had crossed into the host, but such an event was considered a rare anomaly except in very simple organisms. Bacterial DNA is very conspicuous in its structure, so if scientists sequencing a nematode genome, for example, come across bacterial DNA, they would likely discard it, reasonably assuming that it was merely contamination—perhaps a bit of bacteria in the gut of the animal, or on its skin.
But those genes may not be contamination. They may very well be in the host’s own genome. This is exactly what happened with the original sequencing of the genome of the anannassae fruitfly—the huge Wolbachia insert was discarded from the final assembly, despite the fact that it is part of the fly’s genome.
In the early days of the Human Genome Project, some studies appeared to show bacterial DNA residing in our own genome, but those were shown indeed to be caused by contamination. Wolbachia is not known to infect any vertebrates such as humans.
“Such transfers have happened before in the distant past” notes Werren. “In our very own cells and those of nearly all plants and animals are mitochondria, special structures responsible for generating most of our cells’ supply of chemical energy. These were once bacteria that lived inside cells, much like Wolbachia does today. Mitochondria still retain their own, albeit tiny, DNA, and most of the genes moved into the nucleus in the very distant past. Like wolbachia, they have passively exchanged DNA with their host cells. It’s possible wolbachia may follow in the path of mitochondria, eventually becoming a necessary and useful part of a cell.
“In a way, wolbachia could be the next mitochondria,” says Werren. “A hundred million years from now, everyone may have a wolbachia organelle.”
“Well, not us,” he laughs. “We’ll be long gone, but wolbachia will still be around.”
Media Contact
More Information:
http://www.rochester.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
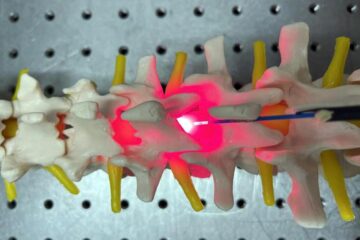
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
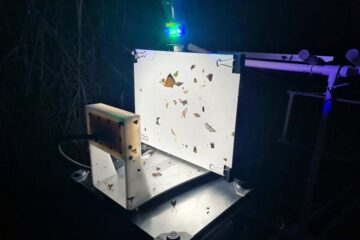

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….