And in the beginning was RNA

And now, while studying the mechanism behind RNAi researchers at Oxford and Helsinki University have discovered that the functional core of a key enzyme (enzymes are proteins which promote biochemical reactions in the body) involved in the formation of RNAi molecules is striking similar to an enzyme involved in gene expression.
The research, published in the journal “Public Library of Science Biology”, supports the idea that the two enzymes have a common ancestor and gives weight to the theory that life started as self-replicating RNA molecules in a RNA world (as opposed to the present world where molecules of DNA are the basis of life).
In fact, we live in a DNA world, as genes are segments of DNA, and it is the information contained in the genes of an organism that, when translated into proteins, makes up the blueprint for the body structure and function. This process, the expression of genes into proteins, is comprised of two steps: the first by which genetic information in DNA is converted into RNA and the second which is the synthesis of proteins based on the information/instructions contained in the newly made RNA (DNA ? RNA ? protein).
But there is a dent in this apparently perfect process. In fact, for a long time scientists have been puzzled why approximately 32% of the human genome/DNA, although transformed into RNA, does not lead to protein production (DNA ? RNA ? no protein)? So why would this huge amount of “junk” RNA keep being formed instead of being eliminated during evolution? After all, a basic rule of life is that any reaction that costs energy and is not advantageous for the individual must be eliminated. What recent research unveiled is that RNA is a much more multifaceted molecule than previously thought, and some of that “junk” RNA actually plays an important role in gene regulation. One such example is RNAi, a RNA that is capable of blocking the activity of specific genes.
And it was while studying the mechanisms behind RNAi, that Paula S. Salgado, Jonathan M. Grimes and colleagues discovered that the functional core of an enzyme involved in the formation of short RNAi molecules from other RNA molecules (RNA? RNA), was remarkably similar to the one that mediates the formation of RNA from DNA (DNA ? RNA) during the first step of gene expression. This striking similarity suggested a common ancestor and further analysis seemed to indicate that the enzyme involved in the RNAi process had appeared before and so would probably be more similar structurally to the common ancestor.
These results support the idea of life starting in a (RNA) world where self-replicating (RNA?RNA) multifunctional RNA molecules evolved (as well as the enzymes mediating the process) into the present situation where genetic information is contained instead on DNA. In fact, although RNA is chemically similar to DNA it has, as the “originater” of life, two major advantages over the latter molecule: 1- it is easily synthesised from non-complex blocks so it had higher possibility of occurring spontaneously and 2 – it is easy to imagine that it could evolve into DNA, which by being a much more stable molecule would then take over. Furthermore, the idea of a primitive RNA world, if proved, could solve one of biggest conundrums on the origin of life: if life needs both DNA as a source of genetic information and proteins to drive life’s chemical reactions how could have one appeared first without the other? Some scientists believe that the answer lies in this ancient RNA molecule which was capable of supporting life reactions and also contained life’s genetic blueprint and whose existence seems to be consistent with the findings of Salgado, Grimes and colleagues.
In this way, Salgado’s work sheds light not only on the mechanism behind this extremely interesting and important process that is RNAi, but can also help to understand better how life began on earth.
Piece researched and written by:
Catarina Amorim ( catarina.amorim@linacre.ox.ac.uk)
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
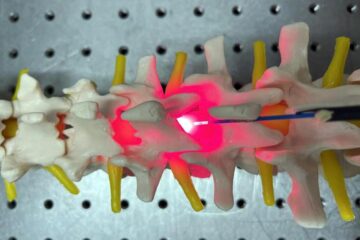
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
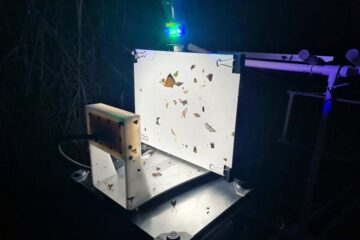

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….