Scientists’ cell discovery unearths evolutionary clues

Researchers, headed by biochemist Professor Pauline Schaap, of the University of Dundee, and evolutionary biologist Professor Sandie Baldauf, of the University of York, have produced the first molecular ‘dictionary’ of the 100 or so known species of social amoeba.
Using this family tree, they have devised a model system to establish how single cell organisms communicate and interact to create multi-cellular structures in response to changing environmental conditions. Previously, there was almost no molecular data for social amoeba – Dictyostelia – which are a hugely diverse and ancient group.
Social amoebas are a group of organisms with a genetic diversity that is greater than that of fungi and similar to that of all animals. They offer an excellent experimental system for studying aspects of evolution and communication that are not easy to study in more complex multi-cellular organisms.
The York and Dundee teams have worked with field biologists in Germany, the US and Japan, and their research is published today (Friday 27th October 2006) in the prestigious international journal Science.
“This provides a starting point in allowing us to examine what happens at the molecular level as species evolve and mutate,” said Professor Schaap, of the Division of Cell and Developmental Biology in the College of Life Sciences at Dundee.
“The availability of a family tree allows us to reconstruct the evolution of the signalling mechanisms that generate multicellularity. It also provides a powerful tool to identify core ancestral processes that regulate the most basic aspects of development.”
Professor Baldauf, of the Department of Biology at York, said: “We have investigated the evolution of plants and animals for a very long time but our whole eco-system depends on single cell organisms. If we want to look at the fundamentals of life we have to look at single cell organisms.
“Amoebas are some of the closest single cell relatives of animals so understanding how they work and evolve is important because it helps us to understand how animals evolve. We have developed a new model system for the study of the evolution of forms.
“We have written the dictionary. Now we know what the words are — but we still have to construct the sentences.”
The research teams were able to build the family tree by amplifying and comparing highly conserved genes from all known species of social amoeba.
The existing family tree of the social amoeba was based on how the multicellular structures of each species look on the outside. However, this tree was completely uprooted by the molecular data gathered by the researchers in Dundee and York.
By plotting all existing information of the amoebas’ cellular and multi-cellular shapes and behaviour to the molecular tree, it appeared that increased cell specialization and organism size is a major trend in the evolution of social amoeba.
Professor Schaap and her team are now working to establish how the regulation and function of genes with important roles in development was altered and elaborated during the course of evolution to generate novel cell-types and morphological features.
The next step for Professor Baldauf and her team will be to investigate the origin of these amoebas, and also to search for new species and to establish their position on the family tree. Meanwhile, a number of research projects, including teams in the USA and Germany, have won sponsorship to sequence the genomes of social amoeba species identified by the work in York and Dundee.
The Dundee-York project was funded under the Biotechnology and Biological Sciences Research Council (BBSRC) CODE (COmparative DEvelopment) initiative and took four years to complete.
Media Contact
More Information:
http://www.dundee.ac.ukAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Why getting in touch with our ‘gerbil brain’ could help machines listen better
Macquarie University researchers have debunked a 75-year-old theory about how humans determine where sounds are coming from, and it could unlock the secret to creating a next generation of more…

Attosecond core-level spectroscopy reveals real-time molecular dynamics
Chemical reactions are complex mechanisms. Many different dynamical processes are involved, affecting both the electrons and the nucleus of the present atoms. Very often the strongly coupled electron and nuclear…
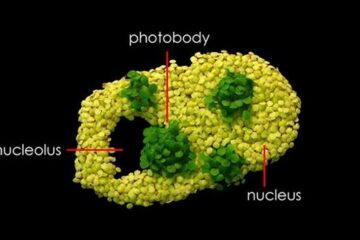
Free-forming organelles help plants adapt to climate change
Scientists uncover how plants “see” shades of light, temperature. Plants’ ability to sense light and temperature, and their ability to adapt to climate change, hinges on free-forming structures in their…