How Purple Corn and RNA Break Genetic Laws

The new research indicates that an additional molecule, DNA's little cousin RNA, is needed for the intriguing gene interactions known as paramutation. Paramutation doesn't follow the laws of classical Mendelian genetics.
“Paramutation is this incredibly interesting, tantalizing violation of Mendel's laws,” said senior author Vicki L. Chandler, director of BIO5 Institute at The University of Arizona in Tucson. “It's been known to exist for 50 years, but nobody understood the underlying mechanism.”
Classical genetics states that when offspring inherit genes from their parents, the genes function in the children the same way the genes functioned in the parent.
When paramutation occurs, one version of the parent's gene orders the other to act differently in the next generation. The gene functions differently in the offspring, even though its DNA is identical to the parent's version.
It happens even when the kids don't inherit the bossy version of the gene. The phenomenon was originally found in corn and has since been found in other organisms, including mammals.
“In previous work we identified a gene that is absolutely required for paramutation to happen,” said Chandler, a UA Regents' Professor of plant sciences and of molecular and cellular biology. “Now we've figured out what that gene does, and it's exciting because it suggests a mechanism for how this process works.”
Chandler's work is the first to point out that an enzyme known as an RNA-dependent RNA polymerase is needed for paramutation.
Corn, also known as maize, is the most economically important crop plant in the United States. Better understanding of plant genetics will help breeders develop improved strains of crops.
Understanding paramutation and similar non-Mendelian genetic phenomena also has implications for human health. For some human diseases, a genetic component is known to exist but has been hard to decipher. Non-Mendelian effects may be at work in those diseases.
“Gene interactions in parents that change the way a gene functions in the progeny are going to contribute to very unexpected inheritance patterns that complicate identifying genes involved in human disease,” said Chandler, who holds the Carl E. and Patricia Weiler Endowed Chair for Excellence in Agriculture and Life Sciences at UA.
Chandler and her colleagues will publish their new findings in the July 20 issue of the journal Nature. The article's title and a complete list of authors and their affiliations is at the end of the release. The National Science Foundation, the National Institutes of Health and the Howard Hughes Medical Institute funded the research.
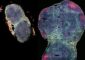
The Chandler lab investigated a gene called b1 that controls whether a corn plant has a purple or green stalk. A plant has two copies of each gene, one from each parent.
One version, or allele, of the gene codes for a purple pigment. Generally, plants need just one copy of that allele, known as B-Intense or B-I, to be the color purple.
But whether a B-I-carrying plant is actually purple depends on the company B-I keeps. If the plant's other b1 allele is the “paramutagenic” B' variety, the B-I allele is silenced. The resulting plant is mostly green.
And although B-I's DNA doesn't change, in subsequent generations the silenced B-I allele behaves as if it had mutated — the B-I-carrying progeny are mostly green, rather than being deep purple.
“It cannot revert — it's a one-way street,” said co-author Lyudmila Sidorenko, an assistant research scientist in Chandler's lab.
Chandler and her colleagues wanted to know how the B' allele changed B-I's behavior without actually changing B-I's DNA. They already knew that paramutation required normal versions of the mediator of paramutation 1 (mop1) gene.
Plants with normal mop1 genes and one B-I allele and one B' allele turned out as expected — mostly green.
However, B-I/B' plants with two mutant mop1 genes were deep purple — they looked as if the purple-suppressing B' allele wasn't present. This demonstrated that normal mop1 was necessary for the B' allele to silence B-I.
The scientists mapped mop1's location on one of the corn's chromosomes and cloned the gene. The mop1 gene makes an enzyme called RNA-dependent RNA polymerase (RDRP). Mutant mop1 genes can't produce the enzyme.
The team had previously suspected a role for RNA, best known for mediating the transfer of information from DNA to a cell's protein-making machinery. This new result provides strong evidence that RNA is indeed involved.
The researchers hypothesize that mop1 amplifies the RNA signals coming from a key region of the B-I and B' allele. That key region is a particular DNA sequence that is repeated seven times.
The researchers hypothesize that those many RNA molecules silence the B-I and B' alleles.
Chandler said, “It's exciting because it's a new role for RNA.”
The researchers' next step is figuring out exactly how RNA suppresses the function of the b1 gene and how those cease-and-desist orders are faithfully transmitted to progeny in the absence of changes in the DNA.
Chandler's co-authors on the article, “An RNA-dependent RNA polymerase is required for paramutation in maize,” are Mary Alleman of Duquesne University in Pittsburgh; Lyudmila Sidorenko, Karen McGinnis and Kristin Sikkink of UA; Vishwas Seshadri, now of Biologics Development Center Developing Businesses in Andhra Pradesh, India; Jane E. Dorweiler, now of Marquette University in Milwaukee; and Joshua White, now of the University of Texas at Austin.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
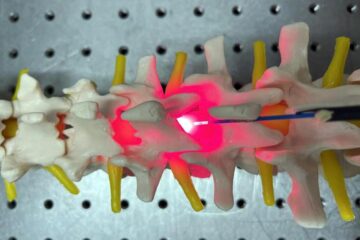
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
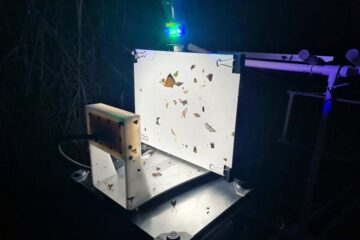

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….