MicroRNA tweaks protein that controls early heart development

Researchers at UT Southwestern Medical Center have discovered how a small molecule of RNA called microRNA – a chemical cousin of DNA – helps fine tune the production of a key protein involved in the early development of heart muscle.
The findings, available in the online edition of the journal Nature, may aid scientists in their understanding of how a progenitor cell, or stem cell, decides to become a heart cell, as well as offer researchers a way to predict how other microRNAs in the body control the production of important proteins. The discoveries could provide clues important to understanding both stem cell biology and congenital heart disease.
In order for cells to produce the proteins that carry out all of life’s functions, the information contained in genes is first copied by special enzymes into messenger RNA, or mRNA. Information in mRNA then is used to make a particular protein.
Scientists believe microRNAs seek out and bind to mRNA, fine tuning the amount of protein that mRNAs manufacture. In some cases, microRNAs shut down protein production altogether.
The UT Southwestern researchers discovered that a microRNA called miR-1 targets the mRNA of the gene Hand2, a key regulator of heart formation. The microRNA turns off production of the Hand2 protein at precisely the right time to allow the proper development of heart muscle.
“We think that Hand2 is necessary in the early stages of embryonic development to allow proliferation and expansion of a pool of muscle progenitor cells that can eventually develop into the heart,” said Dr. Deepak Srivastava, senior author of the paper. “But at some point production of Hand2 needs to be shut off so the cells can go on to the next stage in their development and differentiate into heart muscle cells. We identified Hand2 as the target for this particular microRNA.”
Dr. Srivastava is a former professor of pediatrics and molecular biology at UT Southwestern, where he and his colleagues performed the Nature research. He currently is director of the Gladstone Institute of Cardiovascular Disease and professor of pediatrics at the University of California, San Francisco.
Dr. Srivastava said that if the microRNA is not functioning properly, heart development could be affected in many ways, including not having enough cells or having too many cells in certain locations.
“There are a variety of things that are critical to any organ’s development,” he said. “The Hand2 protein is a master regulator, and in its absence, you don’t get any expansion of the heart ventricle at all. The finding that this microRNA controls Hand2, and probably several other proteins, is very significant.”
The UT Southwestern research team is currently screening human patients with congenital heart disease for mutations in the gene miR-1 to determine what health effects such a mutation might cause. They also are studying mice and fruit flies lacking miR-1.
Dr. Srivastava said the field of microRNA studies has only recently begun to blossom. One of the key challenges is to determine which messenger RNA any given microRNA will target. Hundreds of genes are known to produce microRNA, but in vertebrates there are only three or four known targets for those hundreds.
“We are learning that microRNAs are a common mechanism through which a cell regulates itself at various stages, both during development and later in life,” he said. “This is a rapid way to regulate protein levels. You can imagine a pool of messenger RNA ready to make a protein, and by virtue of a microRNA, that protein synthesis can immediately be shut off and turned back on based on a cell’s environment or its needs at the time.”
Dr. Yong Zhao, a postdoctoral researcher in Dr. Srivastava’s lab at UT Southwestern, developed a new method to predict targets for vertebrate microRNAs based on the genetic sequence of microRNA genes and the accessibility of the target mRNA. Dr. Zhao analyzed all the known microRNA targets in worms and fruit flies to determine what they had in common, hoping to find clues to help predict unknown targets in mammals. He then used those criteria to search the entire mouse genome for potential microRNA targets.
“If this method of predicting targets turns out to be correct and specific, I think it will go a long way to opening the field more broadly, providing scientists who study microRNAs with an easier way to really figure out what they do,” Dr. Srivastava said.
In addition to Dr. Srivastava and Dr. Zhao, UT Southwestern postdoctoral researcher in pediatrics Dr. Eva Samal also contributed to the research.
Media Contact
More Information:
http://www.utsouthwestern.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Cruise Ship as Data Collector
New Approaches in Ocean Observation… Scientific research – not only confined to dedicated research vessels but also from non-scientific vessels and marine infrastructures. This is one of the ideas promoted…
Experiment opens door for millions of qubits on one chip
Researchers from the University of Basel and the NCCR SPIN have achieved the first controllable interaction between two hole spin qubits in a conventional silicon transistor. The breakthrough opens up…
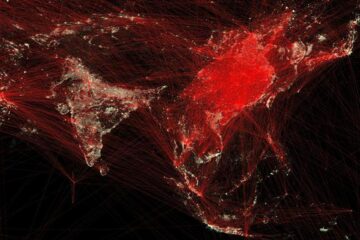
Economies take off with new airports
A global study by an SUTD researcher in collaboration with scientists from Japan explores the economic benefits of airport investment in emerging economies using nighttime satellite imagery. Be it for…





















