UC researchers find new ways to regulate genes, reduce heart damage

Following a heart attack, cells die, causing lasting damage to the heart.
Keith Jones, PhD, a researcher in the department of pharmacology and cell biophysics, and colleagues are trying to reduce post-heart attack damage by studying the way cells die in the heart—a process controlled by transcription factors.
Transcription factors are proteins that bind to specific parts of DNA and are part of a system that controls the transfer of genetic information from DNA to RNA and then to protein. Transfer of genetic information also plays a role in controlling the cycle of cells—from cell growth to cell death.
“We call it 'gene regulatory therapy,'” says Jones.
So far, studies have identified the role for an important group of interacting transcription factors and the genes they regulate to determine whether cells in the heart survive or die after blood flow restriction occurs.
Often, scientists use virus-like mechanisms to transfer DNA and other nucleic acids inside the body.
The “virus” takes over other healthy cells by injecting them with its DNA. The cells, then transformed, begin reproducing the virus' DNA. Eventually they swell and burst, sending multiple replicas of the virus out to conquer other cells and repeat the process.
Now, UC researchers are further investigating new, non-viral delivery mechanisms for this transfer of DNA.
“We can use non-viral delivery vehicles to transfer nucleic acids, including transcription factor decoys, to repress activation of specific transcription factors in the heart,” Jones says, adding that the researchers have made this successfully work within live animal models. “This means we can block the activity of most transcription factors in the heart without having to make genetically engineered mice.”
Jones will be presenting these results at the International Society for Heart Research in Cincinnati, June 17-20.
He says this delivery mechanism involves flooding the cells with “decoys” which trick the transcription factors into binding to the decoys rather than to target genes, preventing them from activating those genes.
“We can use this technology to identify the target genes and then investigate the action of these genes in the biological process,” Jones says.
He says that this delivery has limitations and advantages.
“It can be used to block a factor at any point in time and is reversible,” he says. “However, right now, a specific delivery route must be used to target the tissue or cell.”
Jones and other researchers are hoping that this new technology will allow them to directly address the effects of gene regulation in disease, as opposed to using classical drugs that treat symptoms or have significant adverse outcomes.
“So far, this seems to cause no adverse effects in animals,” he says. “We are hopeful and are working toward pre-clinical studies.”
Media Contact
More Information:
http://www.uc.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
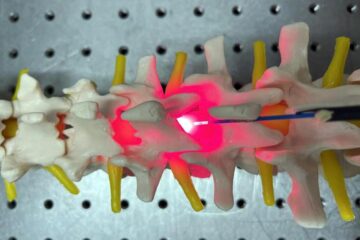
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
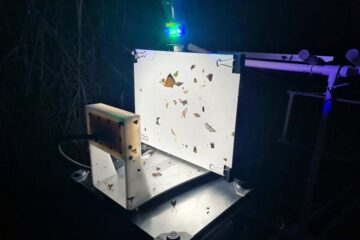

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….