Novel gene-finding approach yields a new gene linked to key heart attack risk factor

Scientists have discovered a previously unrecognized gene variation that makes humans have healthier blood lipid levels and reduced risk of heart attacks — a finding that opens the door to using this knowledge in testing or treatment of high cholesterol and other lipid disorders.
But even more significant is how they found the gene, which had been hiding in plain sight in previous hunts for genes that influence cardiovascular risk.
This region of DNA where it was found had been implicated as being important in controlling blood lipid levels in a report from several members of the same research team in 2008. But although this DNA region had many genes, none of them had any obvious link to blood lipid levels. The promise of an entirely new lipid-related gene took another six years and a new approach to find.
In a new paper in Nature Genetics, a team from the University of Michigan and the Norwegian University of Science and Technology report that they zeroed in on the gene in an entirely new way.
The team scanned the genetic information available from a biobank of thousands of Norwegians, focusing on variations in genes that change the way proteins function. Most of what they found turned out to be already known to affect cholesterol levels and other blood lipids.
But one gene, dubbed TM6SF2, wasn't on the radar at all. In a minority of the Norwegians who carried a particular change in the gene, blood lipid levels were much healthier and they had a lower rate of heart attack. And when the researchers boosted or suppressed the gene in mice, they saw the same effect on the animals' blood lipid levels.
“Cardiovascular disease presents such a huge impact on people's lives that we should leave no stone unturned in the search for the genes that cause heart attack,” says Cristen Willer, Ph.D., the senior author of the paper and an assistant professor of Internal Medicine, Human Genetics and Computational Medicine & Bioinformatics at the U-M Medical School.
“While genetic studies that focused on common variations may explain as much as 30 percent of the genetic component of lipid disorders, we still don't know where the rest of the genetic risk comes from,” Willer adds. “This approach of focusing on protein-changing variation may help us zero in on new genes faster.”
Willer and Kristian Hveem of the Norwegian University of Science and Technology led the team that published the new result. Intriguingly, Willer and colleagues suggest the same gene may also be involved in regulating lipid levels in the liver — a finding confirmed by the observations of a team led by Jonathan Cohen and Helen Hobbs, who propose a role for the gene in liver disease in the same issue of Nature Genetics.
Hveem, a gastroenterologist, says that “more research into the exact function of this protein will be needed to understand the role it plays in these two diseases, and whether it can be targeted with new drug therapies to reduce risk — or treat — one or both diseases.”
The success of the scientific experiment was due to efficient screening of thousands of Norwegian samples and clinical information amassed over a 30-year period by the The Nord-Trøndelag Health Study (HUNT) and the Tromsø Study.
The HUNT Biobank was selected as the “European Research Biobank of the Year” in 2013. Hveem, its managing director, says, “We knew this day would come, when we would see the scientific success stories of decades of labor and tens of thousands of participants donating samples, and it is very rewarding.”
Lead author on the study, Oddgeir Holmen of Norwegian University of Science and Technology, adds, “These are exciting times for disease genetics. The combination of large population-based studies and the rapid development in genotyping technologies will probably help us understand a great deal more about cardiovascular disease, and other diseases, in the next few years.”
Funding:
The HUNT Study is a collaboration between the HUNT Research Centre (Faculty of Medicine, Norwegian University of Science and Technology NTNU), the Nord-Trøndelag County Council, the Central Norway Health Authority and the Norwegian Institute of Public Health.
Other funding came from the U.S. National Institutes of Health, including grants HL094535, HL117626, HL109946, HL068878 and HL117491 from the National Heart, Lung and Blood Institute, DK062370 from the National Institute of Diabetes and Digestive and Kidney Diseases, and HG007022 from the National Human Genome Research Institute.
Authors: In addition to Willer, Hveem and Holmen, the authors include He Zhang, Yanbo Fan, Daniel H. Hovelson, Ellen M Schmidt, Wei Zhou, Yanhong Guo, Ji Zhang, Arnulf Langhammer, Maja-Lisa Løchen, Santhi K. Ganesh, Lars Vatten, Frank Skorpen, Håvard Dalen, Jifeng Zhang, Subramaniam Pennathur, Jin Chen, Carl Platou, Ellisiv B. Mathiesen, Tom Wilsgaard, Inger Njølstad, Michael Boehnke, Y Eugene Chen, and Gonçalo R Abecasis, the director of the Center for Statistical Genetics at the U-M School of Public Health.
Reference: Nature Genetics, 10.1038/ng.2926
Media Contact
More Information:
http://www.umich.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Why getting in touch with our ‘gerbil brain’ could help machines listen better
Macquarie University researchers have debunked a 75-year-old theory about how humans determine where sounds are coming from, and it could unlock the secret to creating a next generation of more…

Attosecond core-level spectroscopy reveals real-time molecular dynamics
Chemical reactions are complex mechanisms. Many different dynamical processes are involved, affecting both the electrons and the nucleus of the present atoms. Very often the strongly coupled electron and nuclear…
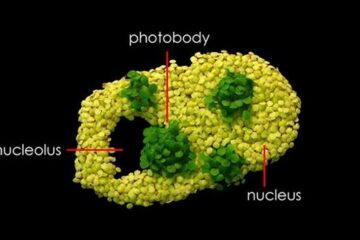
Free-forming organelles help plants adapt to climate change
Scientists uncover how plants “see” shades of light, temperature. Plants’ ability to sense light and temperature, and their ability to adapt to climate change, hinges on free-forming structures in their…