Catching the common cold virus: BYU researchers coming down with the rhinovirus genome

The study is reported in the April issue of the academic journal Molecular Biology and Evolution.
“There are a lot of different approaches to treating the cold, none of which seem to be effective,” said Keith Crandall, professor of biology and co-author of the study. “This is partly because we haven't spent a lot of time studying the virus and its history to see how it's responding to the human immune system and drugs.”
The BYU team studied genomic sequences available online and used computer algorithms to estimate how the rhinovirus is related to other viruses.
According to Nicole Lewis-Rogers, a postdoctoral fellow in the Biology Department and lead author on the study, the rhinovirus is similar to the polio virus, whose vaccine was announced in 1955. But while the polio virus has just three subspecies, the rhinovirus has more than 100 subspecies, which continually evolve.
“These viruses could be under the same constraints and yet change differently,” Lewis-Rogers said. “That's why it is so hard to create a vaccine.”
Through a computer program developed at BYU, Lewis-Rogers' team was able to identify the parts of the virus genome that enable resistance to drugs and the human immune system.
The immune system does a good job of recognizing viral contaminants and getting rid of them, as do new drugs, but the rhinovirus has responded to these defenses by changing its genome so that it is not so easily recognized.
“The virus is evolving solutions against the immune system and drugs,” Crandall said. “The more we can learn about how the virus evolves solutions, the better we can rid the body of these infections.”
Understanding where change occurs in the virus genome will help virologists who work to design drugs that target the rhinovirus.
“If you've got 10,000 bits of information, this narrows it down to a handful,” Lewis-Rogers said. “Here is where you can start looking.”
Lewis-Rogers and Crandall hope scientists will use these insights to build better drugs to combat the virus in the most effective way.
BYU undergraduate Matthew Bendall is also a co-author on the study, which was funded by the USDA. Bendall will next pursue a master's in bioinformatics at BYU.
Media Contact
More Information:
http://www.byu.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
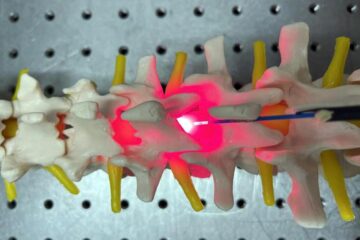
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
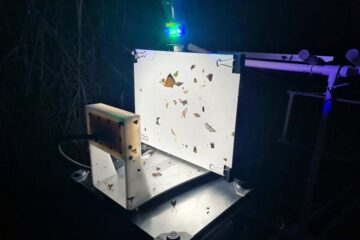

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….