Cloud Computing Method Greatly Increases Gene Analysis

RNA sequencing is used to compare differences in gene expression to identify those genes that switched on or off when, for instance, a particular disease is present. However, sequencing instruments can produce billions of sequences per day, which can be time-consuming and costly to analyze.
The software, known as Myrna, uses “cloud computing,” an Internet-based method of sharing computer resources. Faster, cost-effective analysis of gene expression could be a valuable tool in understanding the genetic causes of disease. The findings are published in the current edition of the journal Genome Biology. The Myrna software is available for free download at http://bowtie-bio.sf.net/myrna.
Cloud computing bundles together the processing power of the individual computers using the Internet. A number of firms with large computing centers including, Amazon and Microsoft, rent unused computers over the Internet for a fee.
“Cloud computing makes economic sense because cloud vendors are very efficient at running and maintaining huge collections of computers. Researchers struggling to keep pace with their sequencing instruments can use the cloud to scale up their analyses while avoiding the headaches associated with building and running their own computer center,” said lead author, Ben Langmead, a research associate in the Bloomberg School’s Department of Biostatistics. “With Myrna, we tried to make it easy for researchers doing RNA sequencing to reap these benefits.”
To test Myrna, Langmead and colleagues Kasper Hansen, PhD, a postdoctoral fellow, and Jeffrey T. Leek, PhD, senior author of the study and assistant professor in the Department of Biostatistics, used the software to process a large collection of publicly available RNA sequencing data. Processing time and storage space were rented from Amazon Web Services. According to the study, Myrna calculated differential expression from 1.1 billion RNA sequencing reads in less than 2 hours at cost of about $66.
“Biological data in many experiments—from brain images to genomic sequences—can now be generated so quickly that it often takes many computers working simultaneously to perform statistical analyses,” said Leeks. “The cloud computing approach we developed for Myrna is one way that statisticians can quickly build different models to find the relevant patterns in sequencing data and connect them to different diseases. Although Myrna is designed to analyze next-generation sequencing reads, the idea of combining cloud computing with statistical modeling may also be useful for other experiments that generate massive amounts of data.”
The researchers were supported by grants from Amazon Web Services, the National Institutes of Health and the Johns Hopkins Bloomberg School of Public Health.
Media Contact
More Information:
http://www.jhsph.eduAll latest news from the category: Information Technology
Here you can find a summary of innovations in the fields of information and data processing and up-to-date developments on IT equipment and hardware.
This area covers topics such as IT services, IT architectures, IT management and telecommunications.
Newest articles
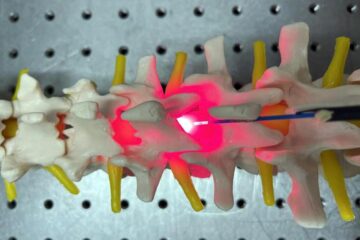
Red light therapy for repairing spinal cord injury passes milestone
Patients with spinal cord injury (SCI) could benefit from a future treatment to repair nerve connections using red and near-infrared light. The method, invented by scientists at the University of…
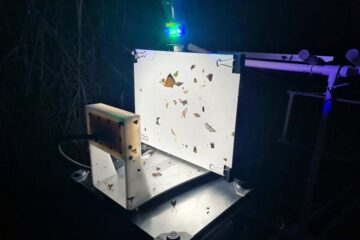

Insect research is revolutionized by technology
New technologies can revolutionise insect research and environmental monitoring. By using DNA, images, sounds and flight patterns analysed by AI, it’s possible to gain new insights into the world of…

X-ray satellite XMM-newton sees ‘space clover’ in a new light
Astronomers have discovered enormous circular radio features of unknown origin around some galaxies. Now, new observations of one dubbed the Cloverleaf suggest it was created by clashing groups of galaxies….