Start codons in DNA may be more numerous than previously thought
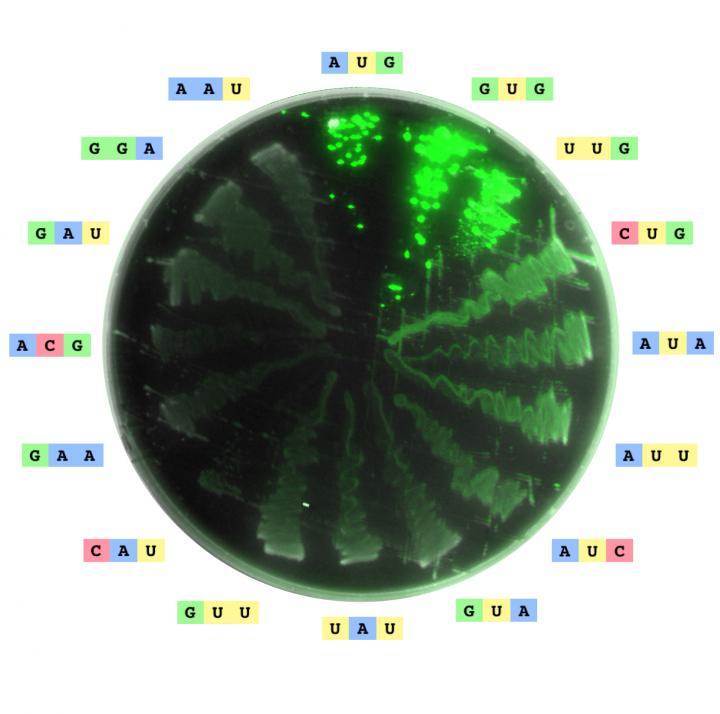
Image of an agar plate streaked with 16 different strains of Escherichia coli, each containing a green fluorescent protein with a different start codon (annotated along the edge of the plate). The 16 codons correspond to the 16 strongest expressing codons. Image is a composite of two super-imposed images from a laser scanner. Credit: Jeff Glasgow/Ariel Hecht/Kelly Irvine/NIST
The findings, to be published on February 21, 2017, in the journal Nucleic Acids Research by scientists in a research collaboration between NIST and Stanford University, demonstrate that there are at least 47 possible start codons, each of which can instruct a cell to begin protein synthesis. It was previously thought that only seven of the 64 possible triplet codons trigger protein synthesis.
“It could be that many potential start codons had remained undiscovered because no one could see them,” said lead author Ariel Hecht, a team member at the Joint Initiative for Metrology in Biology, a research collaboration that includes NIST and Stanford.
Scientists made many of their initial discoveries about DNA and RNA, including start codons, in the 1950s and 1960s. Those ideas have since become enshrined in textbooks around the globe as the modern understanding of the rules of molecular biology.
Genetic code is typically represented via sequences of four letters–A, C, G, and T or U–which correspond to the molecular units known as adenine, cytosine, guanine and thymine (for DNA code) or uracil (for RNA code). Fifty years ago, the best available research tools indicated that there were only a few start codons (with sequences of AUG, GUG and UUG) in most living things. Start codons are important to understand because they mark the beginning of a recipe for translating RNA into specific strings of amino acids (i.e., proteins).
The JIMB team's realization that there might be something amiss in the general understanding of how codons perform began unexpectedly over a round of bagels and coffee. Hecht and his colleagues Jeff Glasgow, Lukmaan Bawazer and Matt Munson were discussing colleague Paul Jaschke's unsuccessful attempt to refactor a virus, phiX174. Refactoring is a kind of re-coding or rearranging used to study genomes and to identify essential genes. phiX174 can be used to infect E. coli cells as a part of such studies.
Jaschke had replaced the start codons of several genes with codons that should not have started translation (AUA and ACG). However, to Jaschke's surprise, he was still detecting the expression of those genes that should have been silenced due to removal.
Hecht pondered what seemed like a rather naïve question: Was Jaschke's experimental result actually wrong? What if the results indicated that codons didn't fit a traditional description of start or not, but instead had varying likelihoods to initiate start translation? To the best of their knowledge, no one had ever systematically explored whether translation could be initiated from all 64 codons. No one had ever proved that you cannot start translation from any codon.
“We kind of all collectively asked ourselves: had anyone ever looked?” said Hecht. A further review of available literature on the topic indicated that the answer was no.
Unlike geneticists working a half-century ago, the JIMB team and others who peer into the inner workings of cells now have far more powerful tools at their disposal, including green fluorescent protein (GFP), a protein adapted from jellyfish, and nanoluciferase, another protein adapted from a deep sea shrimp. Both GFP and nanoluciferase emit light when expressed inside cells and have been optimized within the past decade to produce very strong signals that can be used to probe the cells in depth.
“Ten years ago the tools to make this kind of measurement didn't exist,” Hecht said.
NIST specializes in the process of precision measurement, and the start codon challenge proved irresistible to the JIMB team. The collaboration was formed in 2016 with the goal of advancing biomeasurement science and facilitating the process of discovery by bringing together experts from academia, government labs and industry for collective scientific investigations.
With the use of GFP and nanoluciferase, the team measured translation initiation in the bacteria E. coli from all 64 codons. They were able to detect initiation of protein synthesis from 47 codons.
The implications of the work could be quite profound for our understanding of biology.
“We want to know everything going on inside cells so that we can fully understand life at a molecular scale and have a better chance of partnering with biology to flourish together,” said Stanford professor and JIMB colleague and advisor, Drew Endy. “We thought we knew the rules, but it turns out there's a whole other level we need to learn about. The grammar of DNA might be even more sophisticated than we imagined.”
Still, the JIMB team cautions, this paper is really just the first step, and it is unclear what studies of other organisms will reveal.
“We need to be very careful about extrapolating from these findings or applying them to other organisms without further, deeper research,” said Hecht. He hopes that this paper will encourage or inspire other researchers to explore the topic to find even more answers.
“It could be that all codons could be start codons,” Hecht said. “I think it is just a matter of being able to measure them at the right level.”
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
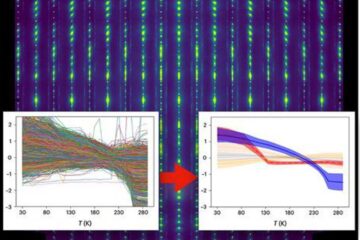
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
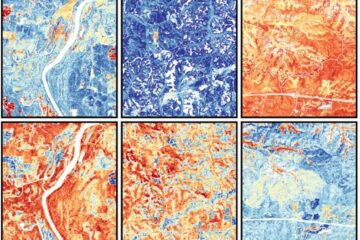

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















