Fission statement
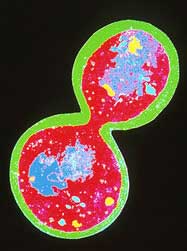
Budding yeast Saccharomyces cerevisiae makes way for a new genome star. <br>© SPL <br>
Alternative yeast joins genome party.
First there was budding yeast (Saccharomyces cerevisiae). Partly responsible for scientists’ survival by fermenting their staples beer and bread, they polished off its DNA sequence back in 1997.
Now the minority fungus of lab culture – fission yeast (Schizosaccharomyces pombe) – is fighting back. This week S. pombe enters the experimental big leagues, with the announcement of its completed genome1.
“For a while they could ignore us completely,” says S. pombe supporter Paul Russell of the Scripps Research Institute in La Jolla, California. “Now they can’t.” With only one yeast genome, “you don’t get the full story”, he argues.
The two yeasts may share a common name, but S. cerevisiae and S. pombe are only distant relatives, having diverged around a billion years ago. “They’ve gone their separate ways,” says evolutionary biologist Russell Doolittle of the University of California, San Diego. As they have developed different ways to solve the basic problems of cell biology, there is still much to learn from the lesser cousin.
Whereas S. cerevisiae cells sprout offspring on their sides, S. pombe divides down the middle, like human cells. The genes and proteins that control this fundamental process in S. pombe were identified by Paul Nurse, the director general of the Imperial Cancer Research Fund in London. His feat won him a share of a 2001 Nobel Prize. What’s more, S. pombe rolls up its DNA into three large chromosomes, again more reminiscent of human cells than S. cerevisiae’s sixteen tiny ones.
Quality over quantity
S. pombe’s genome highlights the dissimilarity between the two yeasts. S. pombe has 4,824 genes – about 1,000 less than its cousin, and fewer even than some bacteria. This is evidence of quality over quantity, says Nurse, who led the sequencing effort.
Despite the yeasts’ differences, “it’s what they share that’s important”, says Nurse. He hopes that comparing the two genomes will reveal exactly which parts are essential for yeast life.
Nurse’s team has already compared the S. pombe genome to another five completed ones – those of budding yeast, the nematode worm Caenorhabditis elegans, the fruit fly (Drosophila), mustard weed (Arabidopsis thaliana) and humans – to find commonalities. All six organisms are ’eukaryotes’ – unlike bacteria, they package up their DNA and protein production into distinct compartments within the cell. The team identified a toolkit of some 60 genes that are essential for organizing and dividing eukaryotic cells.
Armed with the genome, S. pombe researchers hope to wade into its proteome, identifying all its proteins and how they interact. But once again, they’re playing catch-up – budding-yeast researchers are well on their way towards this goal.
Still, S. pombe scientists are used to struggling for recognition. And, despite a slow start and early rivalry, “no one would argue for only one yeast now”, says Russell.
Scientists still harking after S. cervisiae’s brewing power may be won over by S. pombe’s potential. First described by Swiss researcher Lindner in 1893, he isolated it from East African millet beer – and named it after the Swahili word for beer, pombe.
References
- Wood, V. et al. The genome sequence of Schizosaccharomyces pombe. Nature, 415, 871 – 880, (2002).
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Properties of new materials for microchips
… can now be measured well. Reseachers of Delft University of Technology demonstrated measuring performance properties of ultrathin silicon membranes. Making ever smaller and more powerful chips requires new ultrathin…

Floating solar’s potential
… to support sustainable development by addressing climate, water, and energy goals holistically. A new study published this week in Nature Energy raises the potential for floating solar photovoltaics (FPV)…

Skyrmions move at record speeds
… a step towards the computing of the future. An international research team led by scientists from the CNRS1 has discovered that the magnetic nanobubbles2 known as skyrmions can be…





















