A Barcode For Shredding Junk RNA
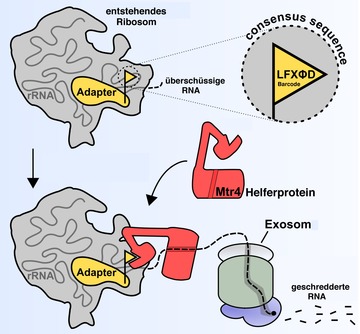
Die schematische Darstellung zeigt, wie das Exosom die überschüssige RNA am entstehenden Ribosom degradiert. Daran beteiligt ist ein Adapterprotein am Prä-Ribosom, das eine Konsensus Erkennungssequenz (Barcode) trägt, an die das Mtr4 Helferprotein binden kann. Dadurch wird das Exosom an die Ziel-RNA dirigiert. Copyright: Universität Heidelberg
A growing, dividing cell uses most of its energy store to make its “protein factories”, the ribosomes. An important player in their “assembly” is the exosome, a molecular shredding machine that breaks down excess ribonucleic acid (RNA).
Researchers working with Prof. Dr. Ed Hurt at the Heidelberg University Biochemistry Center (BZH) have discovered how the exosome identifies its target RNA. The team identified a specific detection signal, comparable to a postal code or bar code that targets the exosome to the remote RNA. The results of the research were published in the journal “Cell”.
According to Prof. Hurt, the production of ribosomes is an extremely complex process that follows a strict blueprint with numerous quality-control checkpoints. The protein factories are made of numerous ribosomal proteins (r-proteins) and ribosomal ribonucleic acid (rRNA).
More than 200 helper proteins, known as ribosome biogenesis factors, are needed in the eukaryotic cells to correctly assemble the r-proteins and the different rRNAs. Three of the total of four different rRNAs are manufactured from a large precursor RNA. They need to be “trimmed” at specific points during the manufacturing process, and the superfluous pieces are discarded. “Because these processes are irreversible, a special check is needed,” explains Ed Hurt.
A substantial portion of this excess ribosomal RNA is broken down by the exosome, a molecular machine consisting of multiple protein subunits. Together they form a structure similar to a barrel, through which the RNA is channelled. The subunit of the exosome that degrades the RNA into its individual components, or nucleotides, sits at the end of this channel. “This process has already been well-described, but we still didn’t understand how the exosome detected its target RNA and distinguished it from the numerous RNA molecules that needed to remain intact,” explains Ed Hurt.
Matthias Thoms and Dr. Emma Thomson of Prof. Hurt’s research team have now been able to identify two ribosome biogenesis factors located near the target RNA on the unfinished ribosome that guide the exosome to its target. Both protein factors act during ribosome assembly, but operate at a different time and position.
Although the two proteins are structured differently, they do share one important characteristic. The Heidelberg researchers discovered both have a short signal sequence similar to a barcode or postal code. Through this detection signal, a helper protein of the exosome is recruited to point to the target RNA. The exosome is then able to start its task and shred the unneeded RNA.
Prof. Hurt’s team of scientists now want to identify other proteins with the described signal sequence to find out how the exosome is able to eliminate such a broad spectrum of different RNA. “The exosome is a universal protein complex that is essential in all cells for RNA homeostasis, that is, the equilibrium between RNA creation and degradation. We assume that the type of target RNA detection we discovered represents a general mechanism for exosome regulation,” explains Prof. Hurt.
“Our findings could also lead to a better molecular understanding of illnesses in which defects were identified in the exosome or their helper proteins.” This would establish the mutations in exosome components that could cause autoimmune diseases or multiple myeloma in humans.
Original publication:
M. Thoms, E. Thomson, J. Baßler, M. Gnädig, S. Griesel and E. Hurt: The Exosome Is Recruited to RNA Substrates Through Specific Adapter Proteins. Cell 162, 1029-1038 (27 August 2015), doi: 10.1016/j.cell.2015.07.060
Contact:
Prof. Dr. Ed Hurt
Heidelberg University Biochemistry Center
Phone +49 6221 54-4781, -4173
ed.hurt@bzh.uni-heidelberg.de
Communications and Marketing
Press Office, phone +49 6221 54-2311
presse@rektorat.uni-heidelberg.de
http://www.uni-heidelberg.de//zentral/bzh/hurt
http://www.bzh.uni-heidelberg.de
Media Contact
More Information:
http://www.uni-heidelberg.deAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…

Rocks with the oldest evidence yet of Earth’s magnetic field
The 3.7 billion-year-old rocks may extend the magnetic field’s age by 200 million years. Geologists at MIT and Oxford University have uncovered ancient rocks in Greenland that bear the oldest…

Decisive breakthrough for battery production
Storing and utilising energy with innovative sulphur-based cathodes. HU research team develops foundations for sustainable battery technology Electric vehicles and portable electronic devices such as laptops and mobile phones are…





















