Worm genes KO'd

Knocking genes out of action allows researchers to learn what genes do by seeing what goes wrong without them. University of Utah biologists pioneered the field. Mario Capecchi won a Nobel Prize for developing knockout mice. Kent Golic found a way to cripple fruit fly genes. Now, biologist Erik Jorgensen and colleagues have devised a procedure for knocking out genes in nematode worms.
“We developed a method that allows us to walk through the worm genome and determine the function of each gene, and thereby infer the function of these genes in humans,” says Jorgensen, a biology professor, senior author of a new study outlining the technique, and an investigator with the Howard Hughes Medical Institute.
The study shows how a transposon or “jumping gene” can be used to delete specific genes from the 1-millimeter-long nematode worm, Caenorhabditis elegans. It was scheduled for online publication Sunday, April 25 in the journal Nature Methods.
“We are trying to understand how genes work and are regulated, and the easiest way to do that is to use a simple organism,” says University of Utah postdoctoral fellow Christian Frøkjær-Jensen, the study's first author. “The amazing thing is that cellular processes in a lowly worm are similar to the biology in humans. We've made it much easier and faster to change the genetic blueprint of a simple worm so we can study and understand how genes are regulated.”
Jorgensen adds: “We want to know what human genes do because they allow us to do all the wonderful things we do – run, speak, live – and understand what goes wrong in genetic diseases and how we can possibly treat them.”
Capecchi, a distinguished professor of human genetics, shared the 2007 Nobel Prize in Physiology or Medicine with two other researchers who independently developed gene targeting in mice in the 1980s. Golic, a professor of biology, published his key papers on targeting or knocking out genes in fruit flies in 2000 and 2002.
“Mario figured out a way to delete genes in mice,” Jorgensen says. “Golic figured out how to do it in fruit flies. And we figured out how to delete genes in worms. There is an institutional excellence in genetics at the University of Utah.”
A Genetic Knife Rather than a Hand Grenade
Genes are on long strands of DNA known as chromosomes. Until now, the main way of altering nematode genes was by using chemicals to induce mutations randomly. The method cannot specify how much DNA should be deleted or exactly where.
“It's like throwing a hand grenade at the genome [genetic blueprint],” Jorgensen says. “Our method is like a surgeon perfectly cutting out a gene.”
He says random mutation doesn't always cripple an entire gene, but just part of it, and can perturb other genes, making it difficult to determine what the targeted gene does.
Random mutagenesis has been able to delete only 5,003 of the 20,160 genes in nematode worms, Jorgensen says. Scientists want to knock out almost all nematode worm genes because “many of the genes in worms are the same as in humans. If we are to understand human genes, it's easier to understand their function in a worm.”
The method developed by Capecchi can hit every gene in mice, crippling them but usually not completely deleting them entirely because mouse genes are large, Jorgensen says. In nematode worms, “we're knocking out the entire gene, but we cannot knock out every individual gene in the worm yet,” although the method should be able to delete 20,043 of 20,160 nematode genes, or 99.4 percent, he adds.
Exploiting a Cell's DNA Repair Mechanism
The new knockout method is named MosDel – for Mos-mediated deletion – because it involves a transposon or jumping gene named Mos1. Jumping genes are pieces of DNA that can jump from one chromosome to another, cutting the DNA where they leave one chromosome and cutting DNA to insert themselves in another.
The Mos1 gene carries the code to make an enzyme named Mos1 transposase. That enzyme does the actual cutting of DNA, Frøkjær-Jensen says.
The transposons in the new study came from fruit flies and were placed into the worm genetic blueprint by French scientists who provided them to the Utah team.
Frøkjær-Jensen says the Utah researchers crippled the jumping genes put into worms “so we can control when and where they hop.”
The Utah biologists used a plasmid – a circular piece of DNA – as a carrier by injecting it with the gene for Mos1 transposase, the “scissors” that cut DNA. Then, a glass needle was used to inject the combination into a worm's gonad.
The transposase-carrying plasmid then cuts out a Mos1 jumping gene adjacent to a gene that researchers want to knock out, leaving a break in the chromosome's DNA.
Cell machinery kicks in to repair the DNA break. Since chromosomes come in pairs, the repair process normally uses the undamaged twin chromosome as a template for repairing the break. But “in this case, we flood the cell with DNA that's similar to where the DNA was broken,” Frøkjær-Jensen says. “We essentially trick the DNA machinery into repairing off a template we supply.”
But the template provided by the biologists lacks the DNA for the gene they want to delete. Thus, the gene is knocked out in the worm's offspring.
DNA is made of “bases” known as G, A, T and C. By “repairing” the broken chromosome, the new method can delete up to 25,000 pairs of those bases. Typical worm genes are 5,000 bases, but range from 700 bases to 30,000 bases.
Frøkjær-Jensen says that to determine how long a segment of worm DNA could be targeted for deletion, the researchers designed repair templates ranging from 1,000 to 50,000 base pairs. They found their method could knock out a gene reliably only if that gene was within 25,000 base pairs of the location where the chromosome is broken. But that still means the method can knock out almost all nematode genes.
Random mutagenesis typically deletes only 300 to 500 base pairs at a time – only part of a gene, Frøkjær-Jensen says.
A Knockout with Benefits
At the same time the DNA repair process is used to delete a targeted worm gene, the scientists also use it to give the mutant worms a “positive selection marker.”
That is because “it's easier to find a moderately sick worm among dying worms than to find one sick worm among a whole lot of healthy worms,” Frøkjær-Jensen says.
So the researchers began with worms lacking a gene named unc-119. Without it, their nerves degenerate and they die. But when the scientists used the DNA repair process to knock out a targeted gene, that process also inserts a working unc-119 gene.
Because worms without unc-119 die, the worms that live are alive because the unc-119 was reinserted at the same time a targeted gene was deleted. So the knockout worms can be identified because they are alive, even if they are ill or abnormal because of whatever targeted gene was knocked out.
The researchers tested the method on 10 genes they targeted for knocking out, including one named dumpy-13. “We knocked out dumpy-13,” Frøkjær-Jensen says. “It made the worms short and fat, so they were easy to distinguish.”
Knocking out genes in worms is advantageous because it is faster and more efficient than other methods. Genes can be knocked out completely, not partially, ensuring that researchers can see what happens without them. Adjacent genes can be deleted, making it easier to learn the combined effects of genes with similar functions.
Frøkjær-Jensen says it might be possible to use the new method to study genes in other animals that serve as models for humans, including flatworms and zebrafish. But the method probably would not work in mammals because deleting a 25,000-base stretch of mammalian DNA probably would be insufficient for deleting their larger genes.
Jorgensen and Frøkjær-Jensen – who also is affiliated with the University of Copenhagen – conducted the study with several co-authors at the University of Utah's Department of Biology: Michael Bastiani, a professor; M. Wayne Davis and Paola Nix, both research assistant professors; Gunther Hollopeter, a postdoctoral fellow; and undergraduates Rachel Lofgren and Michael Prestgard-Duke.
Other co-authors are zoologist Donald Moerman and technician Jon Taylor, both at the University of British Columbia; and Todd Harris, a former University of Utah doctoral student now with Hi-Line Informatics in Livingston, Mont.
University of Utah Public Relations
201 Presidents Circle, Room 308
Salt Lake City, Utah 84112-9017
(801) 581-6773 fax: (801) 585-3350
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
Machine learning algorithm reveals long-theorized glass phase in crystal
Scientists have found evidence of an elusive, glassy phase of matter that emerges when a crystal’s perfect internal pattern is disrupted. X-ray technology and machine learning converge to shed light…
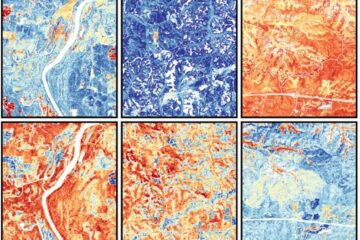

Mapping plant functional diversity from space
HKU ecologists revolutionize ecosystem monitoring with novel field-satellite integration. An international team of researchers, led by Professor Jin WU from the School of Biological Sciences at The University of Hong…

Inverters with constant full load capability
…enable an increase in the performance of electric drives. Overheating components significantly limit the performance of drivetrains in electric vehicles. Inverters in particular are subject to a high thermal load,…





















