When fish swim in the holodeck
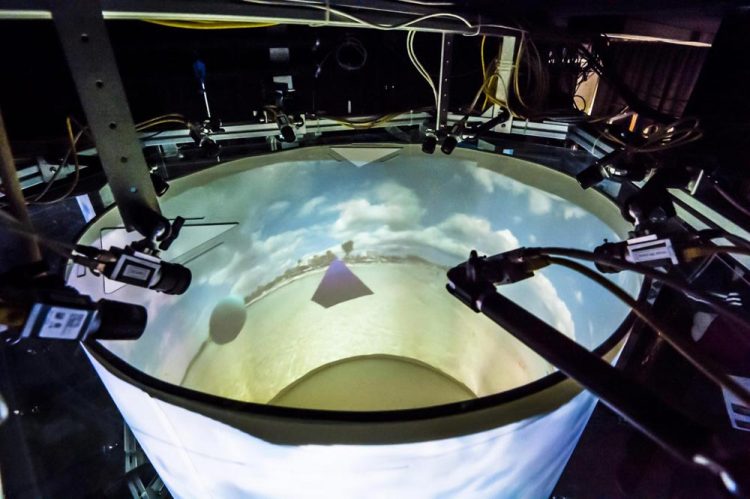

Virtual reality arena for flies. (Copyright: https://strawlab.org/freemovr)
From behavior to brain function
A person sees another person and depending on the context very different interactions can take place. The final outcome after the initial visual experience is a result of complex interactions of neurons in different brain regions- processes that are still very little understood.
To study the neuronal basis underlying behavior, scientists have developed a broad range of techniques, most of which require either the partial or full immobilization of the animal. This restricts sensory input and feedback and ultimately changes the neuronal and behavioral responses. In addition, mimicking natural conditions in a laboratory is difficult.
A three-dimensional, reactive, computer-controlled world for moving animals
The groups of Andrew Straw at the University of Freiburg (a former IMP fellow) and Kristin Tessmar-Raible at the MFPL have now presented a system called “FreemoVR” in Nature Methods, that overcomes most of these hurdles by immersing a freely-moving animal in a reactive, three-dimensional world controlled by a computer.
FreemoVR enables the experimenter to control the animal's visual experience, while maintaining the natural feedback for its tactile senses. To do so, the scientists built behavioral arenas whose walls or floors were computer displays, including arbitrarily shaped projection surfaces. Using computer games technology, the animal could then explore the VR environment in these arenas from its own perspective while it walked, flew or swam.
“We wanted to create a holodeck for animals so that they would experience a reactive, immersive environment under computer control so that we could perform experiments that would reveal how they see objects, the environment, and other animals,” says Andrew Straw, leading in the development of FreemoVR.
Applications of FreemoVR in fish, flies and mice
To validate FreemoVR's ability to elicit naturalistic object responses, the researchers investigated the reaction of freely swimming zebrafish and freely flying flies to a virtual upright post and tested freely walking mice, which showed to be equally afraid of heights in a real and virtual elevated maze.
Using FreemoVR, the teams found previously unnoticed behavioral differences between a wildtype and a mutant zebrafish strain, showing the sensitivity of the system. The scientists further explored the rules that govern social interactions of real zebrafish with virtual ones and found that the prospective leader fish minimizes the risk of losing followers by balancing his internal preference for a swimming direction with the social responsiveness of the subordinate fish.
Future directions
Studying and manipulating behavior in less complex organisms like fish or flies, but also more complex ones like mice and even humans is a popular way among neuroscientists to deduce information about brain function. “I am particularly excited about the possibility to mimic more complex, naturalistic environments and to test more advanced brain functions in medaka and zebrafish. It will help us to better understand brain functions and to what extent we can use these diurnal vertebrates as models for neuropsychological malfunctions”, says MFPL scientist Kristin Tessmar-Raible, who led most of the fish work.
In the future, the different teams hope to use FreemoVR to gain insights into brain function of high-level behaviors like navigation, to better understand causality in collective behavior of social groups and, in the long run, to study the mechanisms of behavior under conditions in which the brain evolved to operate.
###
Publication in Nature Methods: John R Stowers, Maximilian Hofbauer, Renaud Bastien, Johannes Griessner, Peter Higgins, Sarfarazhussain Farooqui, Ruth M Fischer, Karin Nowikovsky, Wulf Haubensak, Iain D Couzin, Kristin Tessmar-Raible, Andrew D Straw. Virtual reality for freely moving animals. Nature Methods, DOI:10.1038/nmeth.4399
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
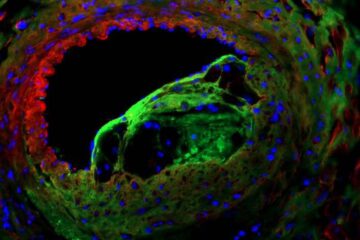
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…

Rocks with the oldest evidence yet of Earth’s magnetic field
The 3.7 billion-year-old rocks may extend the magnetic field’s age by 200 million years. Geologists at MIT and Oxford University have uncovered ancient rocks in Greenland that bear the oldest…

Decisive breakthrough for battery production
Storing and utilising energy with innovative sulphur-based cathodes. HU research team develops foundations for sustainable battery technology Electric vehicles and portable electronic devices such as laptops and mobile phones are…





















