The Splice of Life: Proteins Cooperate to Regulate Gene Splicing
UC San Diego School of Medicine<br>RNAs wound in a knot and bound by hnRNP proteins illustrates the intractable problem of RNA regulation addressed by Huelga et al.<br>
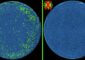
In a step toward deciphering the “splicing code” of the human genome, researchers at the University of California, San Diego School of Medicine have comprehensively analyzed six of the more highly expressed RNA binding proteins collectively known as heterogeneous nuclear ribonucleoparticle (hnRNP) proteins.
This study, published online Feb 16 in Cell Press’ new open-access journal Cell Reports, describes how multiple RNA binding proteins cooperatively control the diversity of proteins in human cells by regulating the alternative splicing of thousands of genes.
In the splicing process, fragments that do not typically code for protein, called introns, are removed from gene transcripts, and the remaining sequences, called exons, are reconnected. The proteins that bind to RNA are important for the control of the splicing process, and the location where they bind dictates which pieces of the RNA are included or excluded in the final gene transcript — in much the same fashion that removing and inserting scenes, or splicing, can alter the plot of a movie.
“By integrating vast amounts of information about these key binding proteins, and making this data widely available, we hope to provide a foundation for building predictive models for splicing and future studies in other cell types such as embryonic stem cells,” said principal investigator Gene Yeo, PhD, assistant professor in the Department of Cellular and Molecular Medicine and the Institute for Genomic Medicine at UC San Diego, and a visiting professor at the Molecular Engineering Laboratory in Singapore. “If we can understand how these proteins work together and affect one another to regulate alternative splicing, it may offer important clues for rational drug design.”
The data sets highlighted in this study – derived from genome-wide methods including custom-designed splicing-sensitive microarrays, RNA sequencing and high-throughput sequencing to identify genome-wide binding sites (CLIP-seq) — map the functional binding sites for six of the major hnRNP proteins in human cells.
“We identified thousands of binding sites and altered splicing events for these hnRNP proteins and discovered that, surprisingly these proteins bind and regulate each other and a whole network of other RNA binding proteins, suggesting that these proteins are important for the homeostasis of the cell,” said first author, NSF fellow Stephanie C. Huelga.
According to the UCSD researchers, the genes specifically targeted by the RNA binding proteins in this study are also often implicated in cancer. Yeo added that of the thousands of genomic mutations that appear in cancer, a vast majority occur in the introns that are removed during splicing; however, intronic regions are where regulatory hnRNP proteins often bind.
“Our findings show an unprecedented degree of complexity and compensatory relationships among hnRNP proteins and their splicing targets that likely confer robustness to cells. The orchestration of RNA binding proteins is not only important for the homeostasis of the cell, but – by mapping the location of binding sites and all the regulatory places in a gene – this study could reveal how disruption of the process leads to disease and, perhaps, a way to intervene.”
Additional contributors to the study include Anthony Q. Vu, Justin D. Arnold, Tiffany Y. Liang, Patrick P. Liu and Bernice Y. Yan, UCSD Cellular and Molecular Medicine; John Paul Donohue, Lily Shiue and Manuel Ares, Jr., UC Santa Cruz; Shawn Hoon and Sydney Brenner, A*STAR, Singapore.
The study was funded in part by grants from the National Institutes of Health and the UC San Diego Stem Cell Research Program.
Media Contact
More Information:
http://www.ucsd.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Combatting disruptive ‘noise’ in quantum communication
In a significant milestone for quantum communication technology, an experiment has demonstrated how networks can be leveraged to combat disruptive ‘noise’ in quantum communications. The international effort led by researchers…

Stretchable quantum dot display
Intrinsically stretchable quantum dot-based light-emitting diodes achieved record-breaking performance. A team of South Korean scientists led by Professor KIM Dae-Hyeong of the Center for Nanoparticle Research within the Institute for…

Internet can achieve quantum speed with light saved as sound
Researchers at the University of Copenhagen’s Niels Bohr Institute have developed a new way to create quantum memory: A small drum can store data sent with light in its sonic…





















