Species identification in the water bottle
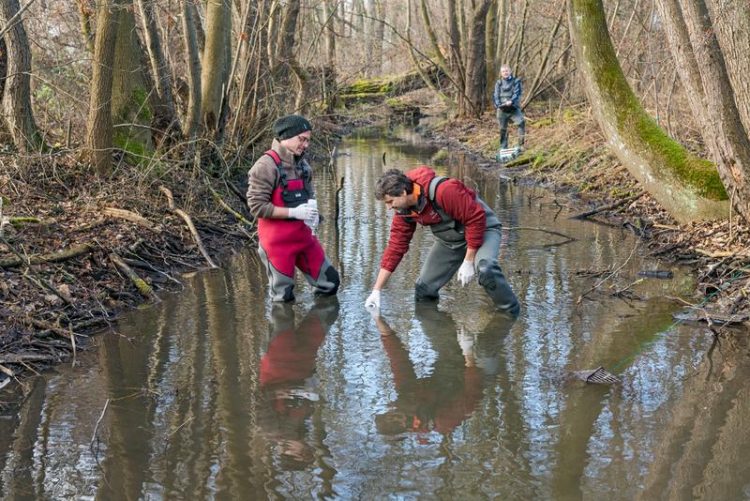

Environmental DNA analysis makes it possible to detect aquatic organisms without having to catch them: Bernhard Stoeckle (right) fetches a water bottle with liquid from a stream. Photo: A. Heddergott/ TUM
DNA that animals release into aquatic environment can be detected using molecular analysis. This detection method is called environmental DNA (eDNA) A simple water sample is sufficient for this technique.
However, this method does not work equally well in all water bodies and is therefore very likely to be influenced by the respective conditions in each body of water. This may include organic and inorganic components in the water or the flow conditions. So far, there is almost no research on how strongly the individual factors affect the analysis procedure.
Dr. Bernhard Stoeckle and Dr. Sebastian Beggel, researchers at the Chair of Aquatic Systems Biology and the Unit of Molecular Zoology (Chair of Zoology) at TUM investigated the influence of a wide range of environmental factors on eDNA analysis in an experiment. The idea for the experiment was based on a previous eDNA study on a native mussel species.
Systematic experimental structure
In a systematic laboratory setup, fish belonging to an invasive species — the round goby (Neogobius melanostomus) — were kept in aquaria in varying densities, under various flow conditions, with and without sediment, and were removed from the water after a defined period of time.
Subsequently, the researchers took water samples at regular intervals over a period of six days in order to be able to evaluate the efficiency of the eDNA analysis over time as well. In addition, the researchers added several substances, which could potentially hinder molecular analysis such as algae, humic substances, and inorganic suspended particles to the water, which are also found in natural ecosystems.
“That’s particularly important, otherwise we wouldn’t be able to apply the findings to eDNA studies in the field,” Bernhard Stoeckle explained. In order to find out which factors have the greatest influence, the researchers then compared the eDNA results of all water samples with each other.
Extent of influence of factors changes over time
On the one hand, the evaluation of the experiment showed that, over its entire duration, the flow conditions, the existence or absence of sediment, and the fish density only had an effect on the analyses in combination with each other. On the other hand, it turned out that the extent of the influence of the factors changed greatly over time.
Of the inhibitors added, organic substances (humic substances) interfered with the analyses the most. They often entirely precluded the successful application of the method. DNA could only be detected in 41 percent of the samples examined. Algae also had a comparable effect, albeit less pronounced. “Our findings clearly demonstrate how important it is that environmental conditions are also taken into account when performing eDNA analyses in order to be able to correctly interpret the findings,” said Bernhard Stoeckle.
Based on the results of the experiment, it can be concluded that specific environmental conditions interfere greatly with environmental DNA experiments, in some cases making it difficult or even impossible to detect species.
Pictures for editorial coverage
Publication:
Stoeckle, BC., Beggel, S., Cerwenka, AF., Motivans, E., Kuehn, R., Geist, J.: A systematic approach to evaluate the influence of environmental conditions on eDNA detection success in aquatic ecosystems, PLoS One 12/2017.
DOI: 10.1371/journal.pone.0189119.
Contact:
Prof. Dr. Jürgen Geist
Technical University of Munich
Chair of Aquatic Systems Biology
Phone: +49/8161/71-3974
Mail: geist@wzw.tum.de
http://fisch.wzw.tum.de
https://www.tum.de/nc/en/about-tum/news/press-releases/detail/article/34398/ article
https://mediatum.ub.tum.de/1427106?id=1427106 pictures
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
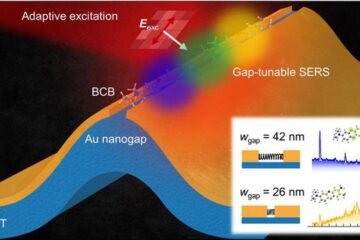
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















