Genome-wide search unearths surprising clues for diabetes and triglycerides

Scientists from the Broad Institute of Harvard and MIT, Lund University and Novartis today announced the discovery of three unsuspected regions of human DNA that contain clear genetic risk factors for type 2 diabetes, and another that is associated with elevated blood triglycerides. The findings stem from the work of the Diabetes Genetics Initiative (DGI), a public-private partnership established in 2004 between Novartis, the Broad Institute of Harvard and MIT, and Lund University, and also reflect a close partnership with two other diabetes research groups. The three groups’ studies, appearing together in the April 26 advance online edition of Science, are among the first to apply a suite of genomic resources to clinical research, including the Human Genome Project, the SNP and HapMap Projects, and genome-scale laboratory and analytical tools.
“For the first time, it is possible to look across the human genome and discover new clues about the root causes of common, devastating diseases that arise from a combination of genes, environment and behavior,” said senior author David Altshuler, a principal investigator of the Diabetes Genetics Initiative, director of the Broad Institute’s Program in Medical and Population Genetics and an associate professor of genetics at Massachusetts General Hospital and Harvard Medical School. “The confirmed genetic contributors we and our collaborators have found open surprising new avenues for disease research, treatment and prevention.”
With the aging of the population and the frequent excesses of modern lifestyles, type 2 diabetes and cardiac risk factors constitute a looming threat to human health, particularly in industrialized nations. Solutions to this burgeoning problem must include new, more effective treatments and the ability to identify “at risk” individuals — each of which requires innovative directions for future research.
The DGI study is one of the first large-scale studies of human genetic variability, aiming to reveal genetic connections to type 2 diabetes and other cardiovascular risk factors such as blood insulin levels, cholesterol levels, blood pressure and body weight. Each of these traits is considered “complex” because it involves a mix of inherited, environmental and behavioral factors.
The scientists’ approach, known as a “genome-wide association study”, involves scanning thousands of individuals’ genomes for single letter changes, called single nucleotide polymorphisms (SNPs). Due to the block-like nature of the human genome, certain SNPs can serve as signposts, highlighting pieces of nearby DNA that may play a causal role in disease.
Using this approach, the DGI team and their collaborators identified and confirmed three novel regions of the genome that influence the risk of type 2 diabetes, as well as a genomic region that is linked with blood triglyceride levels. Perhaps the most intriguing result involves a DNA region that lies far from any known annotated genes. Such genomic “outsiders” would have been incredibly difficult to find by traditional hypothesis-driven approaches.
The other regions linked to diabetes lie near genes with known biochemical functions, but ones never before connected to the disease. Interestingly, the region implicated in triglyceride levels involves a gene that has long been known to play a role in modulating blood glucose.
“This collaboration brings together biologists, geneticists, statisticians and physicians. This interdisciplinary team approach may be the best way to understand complex disorders and quickly turn that to therapeutic advantage,” said Thomas Hughes, Head of Diabetes and Metabolism Research at the Novartis Institutes for BioMedical Research.
Based on initial results, the DGI scientists turned to replicating the most promising findings in independent samples — a critical aspect of the genomic method. The scientists worked together with two other groups that performed similar genomic analyses of type 2 diabetes: the Wellcome Trust Case Control Consortium/U.K. Type 2 Diabetes Genetics Consortium (WTCCC/UKT2D), led by Mark McCarthy and Peter Donnelly of Oxford University and Andrew Hattersley of Peninsula Medical School; and the Finland-United States Investigation of NIDDM Genetics (FUSION) led by Francis Collins of the National Human Genome Research Institute and Michael Boehnke of the University of Michigan.
Notably, the three groups shared their findings in advance of publication to establish priorities for independent follow-up studies. By virtue of their close collaboration, DGI, WTCCC/UKT2D and FUSION researchers identified at least eight clear genetic risk factors for type 2 diabetes, including three that had never before been found, as well as several other probable risk factors that warrant further study.
“This work benefited from unprecedented collaboration among scientists in the diabetes research community, providing the most extensive view yet of the genetic roots of type 2 diabetes. Our own work would not have been possible without the participation of patients in Finland and Sweden, and we are very grateful for their involvement,” said senior author Leif Groop, a principal investigator in the Diabetes Genetics Initiative and a professor in the department of clinical sciences, diabetes and endocrinology at Lund University in Malmo, Sweden. “These findings point at unexpected and new mechanisms contributing to the development of type 2 diabetes, perhaps through the protective mechanisms which allow our insulin-producing cells to cope with the modern lifestyle and environment.”
Genome-wide association studies are the realization of a long-term effort to understand how human genetic variation impacts health. Built on the Human Genome Project, the studies have been made possible in the last year, driven by the recent completion of the HapMap Project and availability of large-scale research tools. Already, scientists from Broad Institute of Harvard and MIT as well as other research organizations worldwide, have used the approach to identify genetic differences that influence a variety of disorders, including Crohn’s disease, obesity, age-related macular degeneration, and prostate cancer.
Media Contact
More Information:
http://www.broad.mit.edu/diabetes/All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles
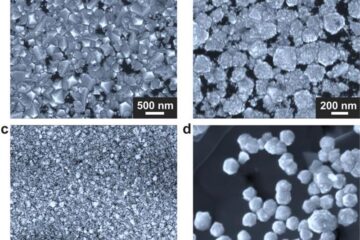

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…





















