Genetic study of Neanderthal DNA reveals early split between humans and Neanderthals

The study helps to explain the evolutionary relationship between Homo sapiens and Neanderthals (Homo neanderthalensis). It also “signifies the dawn of Neanderthal genomics,” wrote the study's authors, who comprise scientists from the Lawrence Berkeley (Calif.) National Laboratory, the U.S. Department of Energy Joint Genome Institute (Walnut Creek, Calif.), the University of Chicago (Ill.) and the Max Planck Institute for Evolutionary Anthropology (Leipzig, Germany).
“Humans went through several key stages of evolution during the last 400,000 years,” said study c-author Jonathan Pritchard, professor of human genetics who led the University of Chicago team that analyzed the sequencing data. “If we can compare humans and Neanderthals genomes, then we can possibly identify what the key genetic changes were during that final stage of human evolution.”
Another author of the Science paper, Svante Pääbo of the Max Planck Institute, sequenced Neanderthal mitochondrial DNA in 1997 and first suggested that Neanderthals did not make a substantial contribution to the modern human gene pool. This new study, headed up by Edward Rubin of the Lawrence Berkeley National Laboratory, reinforces that long-debated theory.
“While unable to definitively conclude that interbreeding between the two species of humans did not occur,” Rubin said, “analysis of the nuclear DNA from the Neanderthal suggests the low likelihood of it having occurred at any appreciable level.”
According to the authors, “If Neanderthal admixture did indeed occur, then [it would] manifest in our data as an abundance of low-frequency derived alleles in Europeans where the derived allele matches Neanderthal. No site in the data set appears to be of this type.”
However, Pritchard said, “We do not exclude the possibility of modest levels of genome admixture.”
Pritchard's team suggests that human and Neanderthal shared a common ancestor about 706,000 years ago, and that the human and the Neanderthal ancestral populations split around 370,000 years ago. (Researchers found some genetic variation between the two species, which the team attributes to the ancestral population.) Both lines co-existed in Europe and western Asia until about 30,000 years ago.
The team used DNA extracted from a 38,000-year-old Neanderthal specimen from Vindija, Croatia. They recovered 65,250 base pairs of the Neanderthal's 3 billion total base pairs and utilized traditional sequencing technologies used for the Human Genome Project as well as the new parallel pyrosequencing method to clone and insert missing fragmented DNA and create a library of Neanderthal DNA.
Unlike the libraries used to sequence the human genome, which contained only human DNA fragments, the Neanderthal DNA library is riddled with contamination from microbes that lived off the nutrients in the Neanderthal remains, as well as contamination from humans handling the specimens.
However, the scientists performed a variety of studies to confirm that the vast majority of the human-like sequence in the library was indeed Neanderthal and not just contamination from human bone collectors and laboratory workers.
The researchers then verified the authenticity of the Neanderthal sequence by comparing it to the human and chimpanzee genomes. This revealed multiple locations where the Neanderthal sequence matched more closely to that of chimpanzee and not human. Using the comparison of the Neanderthal to the human and chimp genomes enabled the investigators to estimate the human-Neanderthal divergence timeline.
The scientists also used data from the HapMap genome project to understand the relationship between modern human diversity and the Neanderthal sequence. Their analysis showed that the Neanderthal sequence could not have come from any modern human population.
The study suggests that Neanderthal and human genomes are greater than 99.5 percent identical, which leaves less than 0.5 percent of the Neanderthal genome that will attract much attention. Many of the biological differences between modern humans and Neanderthals will be encoded at specific sites, which is why the researchers were able to analyze enough data without having to sequence the entire Neanderthal genome.
Media Contact
More Information:
http://www.uchospitals.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
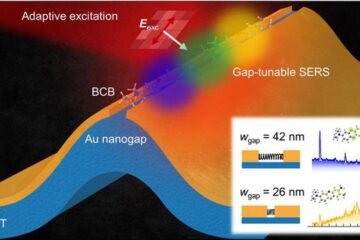
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















