Differences in gene usage dramatically change bacteria’s ’lifestyles’

When and where a bacterium uses its DNA can be as important as what’s in the DNA, according to researchers at Washington University School of Medicine in St. Louis.
Scientists found significant differences in two bacterial organisms’ use of a gene linked to processes that govern a form of antibiotic resistance. The distinction alters the bacteria’s “lifestyles,” or their ability to survive in different environments. Researchers say the finding shows that understanding such changes will likely help development of new treatments for disease-causing microorganisms. “These differences in gene usage are harder to look for, but we’re not going to understand these organisms fully unless we take into account this other dimension,” says senior investigator Eduardo Groisman, Ph.D., professor of molecular microbiology and Howard Hughes Medical Institute investigator.
The study appears the week of Nov. 29 in the online edition of the Proceedings of the National Academy of Sciences and in print on Dec. 7.
One of the bacteria studied, Salmonella enterica, is a leading cause of food poisoning and illness related to animal husbandry. The other, Escherichia coli, can cause illness but more typically plays a beneficial role in the human digestive system. The two are closely related genetically. Less than 20 percent of E. coli’s genes are not found in Salmonella and just over 25 percent of Salmonella’s genes lack counterparts in E. coli.
Groisman’s research had previously focused on how differences in gene content made Salmonella a persistent source of illness. He identified several areas in the bacteria’s DNA known as “pathogenicity islands” — clusters of genes unique to Salmonella that help it cause illness. When complete gene maps for both bacteria became available in recent years, his interests expanded to understanding how the bacteria might use identical genes differently.
Salmonella and E. coli share the gene for an antibiotic resistance regulatory protein called PmrA. By controlling when other proteins are produced, PmrA can make the cell wall more resistant to damage from the antibiotic polymyxin B. The PmrA protein normally activates in response to high iron levels.
In a paper recently published in Genes and Development, Groisman’s lab established that another protein, PmrD, also can activate PmrA in response to low magnesium levels. In the new study, Groisman’s lab discovered that E. coli has a different version of PmrD that is unable to turn on the PmrA protein in response to low magnesium. “We’re not really sure what the significance of low magnesium is, but there are some indications that it may be important to the bacteria’s ability to survive in white blood cells or outside of the host in soil or water,” Groisman says.
When scientists transplanted the Salmonella form of PmrD into E. coli, the bacteria gained the ability to resist polymyxin B in low magnesium environments. Based on data still to be published, Groisman suspects that many other aspects of microbial lifestyle are affected by differences in regulation of identical genes. He notes that the idea of different organisms making altered use of the same genes sprang from recent analyses of the human genome. “Humans not only appear to have far fewer genes than expected, there also seem to be fewer genes that are unique to human DNA than anticipated,” Groisman explains.
In addition to instructions for building proteins, DNA contains stretches of code that affect when genes are turned on and off. As life becomes more complex over the course of evolution, Groisman explains, these regulatory sections appear to take up larger portions of the DNA, allowing genes to be turned on and off in ways that are more intricately responsive to the environment and other factors.
Human DNA, Groisman speculates, may be heavily packed with the factors that allow a more complex, richer use of genes also found in other organisms.
Media Contact
More Information:
http://www.wustl.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

“Nanostitches” enable lighter and tougher composite materials
In research that may lead to next-generation airplanes and spacecraft, MIT engineers used carbon nanotubes to prevent cracking in multilayered composites. To save on fuel and reduce aircraft emissions, engineers…

Trash to treasure
Researchers turn metal waste into catalyst for hydrogen. Scientists have found a way to transform metal waste into a highly efficient catalyst to make hydrogen from water, a discovery that…
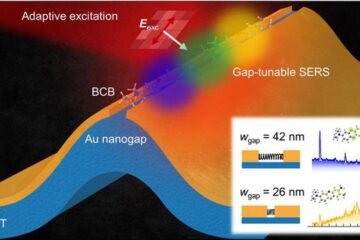
Real-time detection of infectious disease viruses
… by searching for molecular fingerprinting. A research team consisting of Professor Kyoung-Duck Park and Taeyoung Moon and Huitae Joo, PhD candidates, from the Department of Physics at Pohang University…





















