A first glance at global genetic networks
The potential of new technologies to reveal insights into the fundamental structure and function of biological systems continues to grow rapidly –but the ability to interpret and merge these datasets lags behind the ability to collect it. In an effort to overcome these limitations, Sven Bergmann, Jan Ihmels, and Naama Barkai, of the Weizmann Institute of Science in Rehovot, Israel, developed a comparative model that integrates gene expression data from microarrays with genomic sequence information to explore genetic networks.
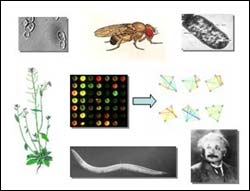
Analyzing the gene expression profiles of six distantly related organisms–bacteria, yeast, plant, nematode, fruitfly, and human–the researchers found that functionally related genes were co-expressed in each species. The most strongly conserved sets of co-expressed genes were associated with core cellular processes or organelles.
Although the regulatory details of individual gene groups varied, the researchers found common ground in the overall landscape of the expression data. The transcription programs exhibit properties typical of dynamically evolving “real-world” networks that are designed to perform in uncertain environments and maintain connections between elements independent of scale. These properties were originally identified in studies of social networks and the World Wide Web, but they aptly describe the real-world challenges of the cell. Studies of dynamically evolving networks show that nodes (i.e., genes and proteins) added at an early stage (much like highly conserved genes) are more likely to develop many connections, acting as a hub. Following these organizational principles, transcription networks would have a relatively small number of highly connected “hub genes”–though a much higher number than one would expect in a random network.
And that is what the authors observed: the networks they constructed from the expression data had the expected number of highly connected hub genes, which tend to be essential and conserved between organisms. Since these highly connected genes are likely to have homologues in other organisms, they can serve as powerful and efficient tools for assigning function to the thousands of uncharacterized sequences found in sequence databases. This model presents a framework to explore the underlying properties that govern the design and function of the cell, and provides important clues–in the form of conserved transcription modules–to the evolutionary building blocks that generate diversity.
All works published in PLoS Biology are open access. Everything is immediately available without cost to anyone, anywhere–to read, download, redistribute, include in databases, and otherwise use–subject only to the condition that the original authorship is properly attributed. Copyright is retained by the author. The Public Library of Science uses the Creative Commons Attribution License.
Media Contact
More Information:
http://www.plosbiology.orgAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















