Carnegie Mellon scientists create unique DNA probe with great potential
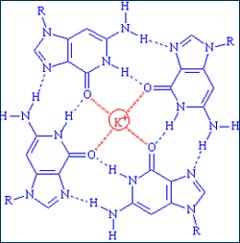
Figure 1. Assembly of four guanine (G) nucleobases into a quadruplex. The potassium ion in the center stabilizes the structure by bonding to the oxygen atoms on each guanine.
A team of investigators at Carnegie Mellon University has formed the first hybrid quadruplex of peptide nucleic acids, or PNAs, with DNA, the genetic code. This result opens new opportunities to study the activity of genetic regions occupied by recently described quadruplex DNA structures, as well as providing a new compound that could be used as a biosensor or to block gene activity associated with diseases such as cancer. The research results, published online, will appear in a forthcoming issue of the Journal of the American Chemical Society.
“PNA2-DNA2 hybrid quadruplexes are extremely stable, suggesting that if we use PNAs to bind with DNA quadruplexes that regulate gene expression, we could prevent disease processes in which these DNA quadruplexes appear to play a role,” says Bruce Armitage, Ph.D., associate professor of chemistry at Carnegie Mellon. “PNAs also could be converted into biosensors by coupling them with fluorescent tags that would allow the PNA to report the presence of a successful hybridization to quadruplex-forming sequences either in the genome or in messenger RNA molecules.”
“In addition to a new high-affinity DNA recognition mechanism and expanding the scope of molecular recognition by PNA, the PNA2-DNA2 hybrid quadruplex is the first example of homologous hybridization,” adds Armitage.
PNA2-DNA2 hybrid quadruplexes, which assume a rod-like structure, could also prove important in nanotechnology applications, such as building a nanoscale bridge to conduct electrical charges or a precisely controlled structural component of a biosynthetic material.
Normally, DNA occurs as the well-known double helix first proposed by James Watson and Francis Crick 50 years ago. Each strand of the helix consists of a backbone of sugar-phosphate groups linked to nucleobases, which occupy the inside of the helix. Nucleobases of one strand bind to nucleobases of a complementary strand, and the two strands wind around one another like a twisted ladder. In recent years, scientists have found that DNA sometimes kinks or bends into unusual structures, such as hairpin turns, triplexes or quadruplexes. In the latter case, four DNA guanine (G) nucleobases align in a wheel-like structure (see Figure 1). DNA-quadruplexes recently have been found in complexes called telomeres, DNA sequences that cap and protect the ends of chromosomes. There, DNA quadruplexes appear to play a role in preventing chromosome ends from fraying when chromosomes duplicate during cell division. DNA quadruplexes also appear to play important roles in regulating the expression of genes that produce antibodies as well as c-MYC, a cancer-causing gene. A quadruplex structure has also been implicated in the messenger RNA derived from FMRP, a gene that has been implicated in fragile X syndrome, the most common form of mental retardation.
At the same time as these discoveries have been made, other research has explored the properties of artificially manufactured PNAs, in which nucleobases are bound to a backbone chain of amino acids, rather than sugar-phosphate groups (see Figure 2). PNAs can be made to target specific DNA sequences and have been shown to bind, or hybridize, with helical DNA according to the pairing rules first proposed by Watson and Crick, namely guanine (G) pairs with cytosine (C), and adenine (A) pairs with thymine (T). The resulting PNA-DNA hybrid duplexes are actually more stable than a duplex consisting of two complementary DNA strands. In addition, PNAs also resist enzymatic breakdown. Given these properties, many investigators consider PNAs highly promising agents for molecular drug design because they have the potential to persist in a cell, target specific genes and regulate gene expression. What distinguishes the recent report from Armitage’s lab is that the PNA used to hybridize with a quadruplex-forming DNA sequence has the homologous (identical) sequence as the DNA, rather than the complementary sequence. DNA quadruplexes can form from four separate G-rich strands, two such strands that fold back on themselves to form hairpin structures, or from a single strand that folds back on itself three times (see Figure 3). In each case, four Gs are positioned properly to allow them to form the G-quartet that stabilizes a quadruplex.
“We reasoned that since DNA quadruplexes can form from multiple DNA strands, then it should be possible to have a PNA strand substitute for a DNA strand, potentially forming a more stable quadruplex. This turned out to be the case,” explains Armitage. In the experiments carried out in Armitage’s lab by graduate student Bhaskar Datta and undergraduate student Christoph Schmitt, the “hybrid quadruplex” assembled from mixing together homologous PNA and DNA strands actually consists of two strands each of PNA and DNA. The strands do not fold back on themselves, leading to formation of an extended four-stranded structure (see Figure 4). Optical methods such as UV-visible, circular dichroism and fluorescence spectroscopy were used to determine the number of strands in the hybrid and their relative orientations. Temperature-dependent experiments revealed that the PNA2-DNA2 hybrid quadruplexes were extremely stable, meaning the strands did not readily separate from one another.
Watson-Crick hybridization between two complementary strands can be applied in principle to any sequence of DNA or RNA, and this forms the basis of numerous clinical diagnostic tests, so-called “gene chips,” and the antisense approach to gene therapy. In contrast, homologous hybridization is restricted to a relatively small number of sequences. Nevertheless, the importance of G-rich sequences such as those successfully targeted by the Carnegie Mellon researchers continues to grow as more and more such sequences are discovered in key regulatory regions of DNA and RNA. In principle, any molecule that can bind to a quadruplex structure with high affinity has the potential for either repressing or stimulating the function of that quadruplex. Before PNAs can be used in these applications, according to Armitage, scientists need to learn more about the factors that control the number of DNA and PNA strands that are incorporated into hybrid quadruplexes. This is where the Armitage group’s current studies are directed.
While the PNA2-DNA2 hybrid quadruplex establishes a new approach to targeting DNA sequence specifically, it is still several steps away from being useful in the clinic. The DNA sequence targeted by homologous PNA is not found in humans, but rather in the telomere of a well-studied model organism, the ciliated protozoan Oxytricha nova. In the laboratory, this sequence has the interesting property of being able to adapt its structure to the surrounding conditions, shifting between a four-stranded quadruplex with all strands extended, as in the PNA2-DNA2 hybrid quadruplex, and a two stranded quadruplex in which the two strands are folded back on themselves to form hairpins. Human DNA has a different sequence, so different PNA strands will need to be synthesized before such experiments can begin. In addition, in vivo, only a single G-rich strand is present, meaning the DNA folds back on itself three times to form an “intramolecular” quadruplex.
“We need to understand the rules for designing a homologous PNA to target an intramolecular quadruplex before we can realistically target these structures inside of cells,” explains Armitage, adding, “We hope that the experiments we are currently performing in the lab will bring us closer to the goal of targeting these fascinating structures in vivo.”
The PNA2-DNA2 hybrid quadruplex might have applications in nanotechnology, where the goal is to build functional assemblies of molecules having nanometer dimensions. The first step in many of the proposed applications of nanotechnology is simply the construction of the device, which will require the precise assembly of individual molecules in two or three dimensions. The well-defined and easily controlled dimensions of DNA, combined with the simple pairing rules for the nucleobases, already has led to the use of double-helical DNA as a construction element for nanostructures, according to Armitage. Any new recognition mode that adds to the structural diversity of DNA offers opportunities for novel nanostructures. In the case of the PNA2-DNA2 hybrid quadruplex, four strands feed into the central quadruplex structure, which can be viewed as a junction element. This is distinct from a Watson-Crick duplex, where only two strands are connected.
“Michael Crichton might not use this in his next book, but the opportunities for building functional nanostructures based on the PNA2-DNA2 hybrid quadruplex are very interesting to us, and we hope to exploit this novel recognition mode,” notes Armitage.
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…





















