Scientists Find That Apes and Monkeys Provide Needed Help in Understanding the Human Genome
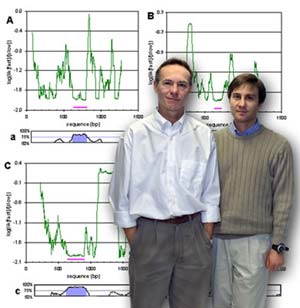
Eddy Rubin (left) along with Dario Boffelli led the development of a technique called phylogenetic shadowing which enables scientists to make meaningful comparisons between the genomes of humans and other primates.
Scientists with the U.S. Department of Energy’s Joint Genome Institute (JGI) and the Lawrence Berkeley National Laboratory (Berkeley Lab) have developed a powerful new technique for deciphering biological information encoded in the human genome. Called “phylogenetic shadowing,” this technique enables scientists to make meaningful comparisons between DNA sequences in the human genome and sequences in the genomes of apes, monkeys, and other non-human primates. With phylogenetic shadowing, scientists can now study biological traits that are unique to members of the primate family.
“Now that the sequence of the human genome has almost been completed the next challenge will be the development of a vocabulary to read and interpret that sequence,” says Edward Rubin, M.D., director of the Joint Genome Institute (JGI) for the U.S. Department of Energy, and Berkeley Lab’s Genomics Division, who led the development of the phylogenetic shadowing technique.
“The ability to compare DNA sequences in the human genome to sequences in non-human primates will enable us in some ways to better understand ourselves than the study of evolutionarily far-distant relatives such as the mouse or the rat,” Rubin adds. “This is important because as valuable as models like the mouse have been, there are many physical and biochemical attributes of humans that only other primates share.”
Using phylogenetic shadowing, Rubin and his colleagues were able to identify the DNA sequences that regulate the activation or “expression” of a gene that is an important indicator of the risk for heart disease and is found only in primates. The results of this research are reported in a paper published the February 28 issues of the journal Science. Co-authoring the paper with Rubin were Dario Boffelli, Dmitriy Ovcharenko, Keith Lewis and Ivan Ovcharenko of Berkeley Lab, plus Jon McAuliffe and Lior Pachter, of the University of California at Berkeley.
Comparative genomics, comparing segments of DNA in the human genome to DNA segments in the genomes of other organisms that have been sequenced, such as the mouse, the puffer fish or the sea squirt, has proven to be an effective means of identifying genes, the DNA sequences that code for proteins, and gene regulatory sequences, the DNA sequences which control when a gene is turned on or off.
“The rationale for comparing the genomes of different animals to identify those sequences that are important is based on the understanding that today’s different animals arose from common ancestors tens of millions of years ago,” Rubin explains. “If segments of the genomes of two different organisms have been conserved (meaning the sequences are the same in both) over the millions of years since those organisms diverged, then the DNA sequences within those segments probably encode important biological functions.”
The search for functional DNA sequences that have been conserved between two different organisms across a large distance in evolution is the classical approach to comparative genomics that has been used to interpret the information in the human genome. In order for this technique to work, the conserved functional sequences have to stand out as distinct from the non-functional sequences which were not conserved. That degree of distinction requires the passage of time — lots of it — in order for mutations and the lack of selection pressures to cause the non-functional sequences in the two genomes to drift apart.
For example, mice and humans last shared a common ancestor about 75 million years ago, plenty of time for the non-functional sequences in their respective genomes to go their separate ways. Only about five-percent of the two genomes are conserved and it has been shown that most of the genes and regulatory sequences that have been discovered lie within these conserved DNA segments. On the other hand, humans and non-human primates shared common ancestors as recently as 6 to 14 million years ago for apes, 25 million years ago for Old World (African) monkeys, and 40 million years ago for New World (South American) monkeys. This is insufficient time for much genetic divergence to have taken place. Consequently, non-human primates have been largely ignored in the effort to interpret the human genome.
“Comparative genomics studies between evolutionarily distant species will readily identify regions of the human genome performing basic biological functions shared with most mammals,” says Rubin. “However, it will invariably miss recent changes in DNA sequence that account for primate-specific biological traits.”
Rubin has likened comparisons between the human and mouse genomes to comparisons between an automobile and a go-cart: “Only the very basic parts and design features are similar.” Whereas, he argues, comparing the human genome to that of a chimp or a baboon, is like comparing a sedan to a station wagon: “Nearly all the parts and design features are almost interchangeable.”
Until now, however, comparing the human genome to that of a chimp or baboon has been a problem since both genomes are so much alike.
As Boffelli, who works with Rubin at both Berkeley Lab and JGI explains, “There is only about a 5-percent difference between the human and the baboon genomes. When you run comparisons between the two, all of the sequences look just about the same. We can’t distinguish function from non-functional sequences.”
Rubin and his colleagues overcame this lack of distinction by comparing segments of the human genome to segments of not one but anywhere from 5 to 15 different genomes of non-human primates, including chimpanzees and gorillas, orangutans, baboons, and Old World and New World monkeys. By sequencing specific segments within each of the genomes of the different primates being analyzed, the researchers found enough small differences from genome to genome in the non-human primates that could be combined to create a phylogenetic “shadow” which could then be compared to the human genome.
“The additive collective sequence differences or divergence of these non-human primates as a group was comparable to that of humans and mice,” Rubin says. “This suggests that deep sequence comparisons of numerous primate species should be sufficient to identify significant regions of conservation that encode functional elements shared by all primates including humans.”
The phylogenetic shadow that Rubin and his colleagues created was distinct enough for them to see the boundaries between exons (protein-coding DNA sequences) and introns (non-coding DNA sequences) for several genes in addition to discovering the regulatory elements for a gene named “apo(a)” which is associated with low-density lipoproteins (LDLs) in the blood stream of humans. An evolutionary new-comer, apo(a) is found in humans, apes, and Old World monkeys but appears to be lacking in nearly all other mammals. Biomedical researchers want to know the regulatory sequences of apo(a) because high blood levels of apo(a) are an important risk predictor for cardiovascular disease. The desire to study apo(a) is the reason Rubin and his research group began the development of their phylogenetic shadowing technique.
“We could not study apo(a) by comparing human DNA sequences to the sequences of evolutionarily distant species as those species don’t have apo(a) so we had to find an alternative method,” Rubin says.
Rubin’s research group at Berkeley Lab has been at the forefront of using transgenic mice and the mouse genome to decipher the human genome and to identify and study important genetic risk factors in the development of human heart disease. He and his group believe that the ability to do comparative genomic studies with non-human primates will prove especially beneficial to human medical research. Their data from this study suggests that sequencing the genomes of as few as four to six primate species in addition to humans may be enough to identify much of the conserved functional DNA sequences in the human genome.
“The argument for sequencing a broad variety of evolutionarily distant species, like the mouse and puffer fish, has been that they would be needed for us to gain a good understanding of the human genome,” Rubin says. “These evolutionarily distant creatures have been incredibly useful but maybe now we should be focusing our effort on sequencing the genomes of not one but several different non-human primates. Their collective sequences will tell us things about the human genome that we will never to able to learn from our more distant relatives in the animal kingdom.”
This research was funded by a grant from the National Heart, Lung, and Blood Institute.
Berkeley Lab is a U.S. Department of Energy national laboratory located in Berkeley, California. It conducts unclassified scientific research and is managed by the University of California. Visit our Website at www.lbl.gov/.
Additional information
Dr. Edward Rubin can be reached at (510)486-5072 or
EMRubin@lbl.gov
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

High-energy-density aqueous battery based on halogen multi-electron transfer
Traditional non-aqueous lithium-ion batteries have a high energy density, but their safety is compromised due to the flammable organic electrolytes they utilize. Aqueous batteries use water as the solvent for…

First-ever combined heart pump and pig kidney transplant
…gives new hope to patient with terminal illness. Surgeons at NYU Langone Health performed the first-ever combined mechanical heart pump and gene-edited pig kidney transplant surgery in a 54-year-old woman…

Biophysics: Testing how well biomarkers work
LMU researchers have developed a method to determine how reliably target proteins can be labeled using super-resolution fluorescence microscopy. Modern microscopy techniques make it possible to examine the inner workings…





















