Biochemists Identify Protease Substrates Important to Bacterial Growth
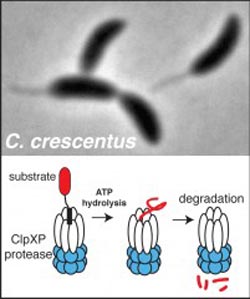
Peter Chien<br> <br>Caulobacter crescentus (above) generates radically different cell types upon division. The ClpXP protease (illustrated below) recognizes and destroys many protein substrates that allow Caulobacter to differentiate into these different cell types. New work identifying scores of new candidate substrates of ClpXP to reveal how protein degradation is critical to cell cycle progression and bacterial development could lead to new antibiotic targets.<br>
As Chien (pronounced Chen) explains, to carry out fundamental life processes such as growing and dividing, cells must orchestrate, in time and location, the production and degradation of hundreds of protein substrates. Even in simple bacteria, protein degradation is critical for making sure these organisms can grow and respond to their environment properly.
Scientists have known that a group of protein machines called energy-dependent proteases are responsible for the majority of this degradation, but what targets these machines recognize and how they do it has been unknown in many cases.
With the new series of experiments in the model bacteria Caulobacter crescentus in the Chien biochemistry and molecular biology laboratory, much more is now understood, he says. “We first generated a protease mutant that could recognize but not destroy its targets, acting as a ‘trap’ for protease substrates. After purifying this trap from living cells, we used mass spectrometry to identify proteins that were caught, finding over a hundred new candidate substrates. These targets covered all aspects of bacterial growth, including DNA replication, transcription and cytoskeletal changes.”
Next, they focused on one of these new targets in detail, a protein called TacA. Caulobacter grow by making two different cell types every time they divide. TacA is responsible for making sure that one of these cell types forms properly.
“We used biochemistry and highly purified proteins to identify what parts of TacA were important for degradation by the ClpXP protease,” Chien says. “We then made mutants of TacA that could not be degraded and found that when we expressed them in bacteria, these cells failed to properly develop into the correct cell types. Because developmental changes are essential for pathogenic bacteria to invade their host, these insights could potentially identify new antibiotic targets.”
The work was funded by a grant from the National Institute of General Medical Sciences at the National Institutes of Health and by UMass Amherst.
Media Contact
More Information:
http://www.umass.eduAll latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…





















