A molecular 'atlas' of animal development
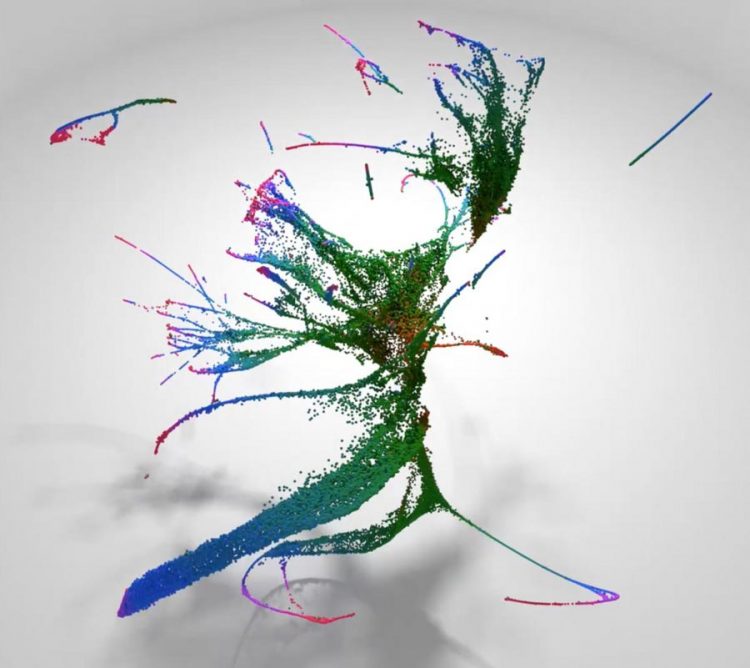

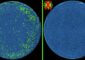
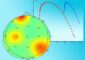
Each cell of a developing nematode worm embryo is catalogued at the molecular level in a new paper out in Science. In this visualization of the dataset, each dot represents a single cell, its color represents the age of the embryo it came from (orange=early, green=mid, blue/red=late), and the dots are arranged so that cells with similar transcriptomes are near each other. Visualized this way, the data form various thin "trajectories" that correspond to tissues and individual cell types. Credit: Cole Trapnell Usage Restrictions: In context of reporting
In a paper in Science this week, Penn researchers report the first detailed molecular characterization of how every cell changes during animal embryonic development. The work, led by the laboratories of Perelman School of Medicine's John I. Murray, the School of Arts and Sciences' Junhyong Kim, and Robert Waterston of the University of Washington (UW), used the latest technology in the emergent field of single cell biology to profile more than 80,000 cells in the embryo of the nematode Caenorhabditis elegans.
“Over the past few years, new single cell genomics methods have revolutionized the study of animal development,” says Murray. “Our study takes advantage of the fact that the C. elegans embryo has a very small number of cells produced by a known and completely reproducible pattern of cell divisions. Using single cell genomics methods, we were able to identify over 87 percent of embryonic cells from gastrulation (when there are about 50 cells present) through the end of embryogenesis.”
C. elegans is an animal that hatches with only 558 cells in its body. In a multicellular organism, every cell is derived by cell division from a single fertilized egg, resulting in a “cell lineage tree” that shows the division history of every cell, and describes their relationships to each other, akin to a genealogy. The Nobel prize winning work of Sydney Brenner, H. Robert Horvitz, and John Sulston worked out the cell lineage tree of C. elegans more than 40 years ago, and showed that every C. elegans animal develops through identical patterns of cell division.
To further elucidate the process of development, the Penn and UW teams characterized what happens at the molecular level by measuring the transcriptome–all the RNAs in a cell–of individual cells during development using a single cell genomics approach. These methods allow scientists to determine which genes are expressed, or turned on, in each of tens or hundreds of thousands of cells and to identify rare cell types based on their expression of similar subsets of the genes. However, it is difficult to know in these studies whether all cell types have been identified, or how the identified cells are related to each other through cell division.
The lead authors, graduate students Jonathan Packer of UW and Qin Zhu of Penn, developed sophisticated data analysis programs and algorithms to trace the changes in the transcriptome to the temporal sequences in the cell lineage tree, revealing detailed dynamics of molecular changes required to generate the full body of C. elegans.
The resulting dataset will be a powerful tool for the thousands of labs that study C. elegans as a model organism and reinforces the limitations of using single cell genomics alone to infer relationships between cells in other species.
“Penn has been one of the pioneers of single cell genomics, which really helped make this work possible,” says Kim.
The investigation helps reveal fundamental mechanisms involved in how cells specialize their function during development. For example, the researchers showed that cells with very different lineage histories can rapidly converge to the same molecular state, such that they can no longer be distinguished. The researchers also found that, during differentiation, some cells undergo strikingly rapid changes in their transcriptomes.
In addition, this work will contribute to applications in regenerative medicine and cellular engineering, such as controlling the cell-differentiation process involved in using patient's own cells for therapy.
###
John Murray is associate professor of genetics in the Perelman School of Medicine at the University of Pennsylvania.
Junhyong Kim is the Patricia M. Williams Term Endowed Professor of Biology in the School of Arts and Sciences at the University of Pennsylvania.
In addition to Murray, Kim, Waterston, Packer, and Zhu, the paper was coauthored by Penn's Priya Sivaramakrishnan, Elicia Preston, Hannah Dueck, Derek Stefanik, and Kai Tan and UW's Chau Huynh and Cole Trapnell.
The study was supported by the National Institutes of Health (grants HG007355, GM072675, GM127093, and HD085201), Commonwealth of Pennsylvania, and Penn Program in Single Cell Biology (co-directed by Kim and James Eberwine, professor of systems pharmacology and translational therapeutics in the Perelman School of Medicine at the University of Pennsylvania).
Media Contact
All latest news from the category: Life Sciences and Chemistry
Articles and reports from the Life Sciences and chemistry area deal with applied and basic research into modern biology, chemistry and human medicine.
Valuable information can be found on a range of life sciences fields including bacteriology, biochemistry, bionics, bioinformatics, biophysics, biotechnology, genetics, geobotany, human biology, marine biology, microbiology, molecular biology, cellular biology, zoology, bioinorganic chemistry, microchemistry and environmental chemistry.
Newest articles

Security vulnerability in browser interface
… allows computer access via graphics card. Researchers at Graz University of Technology were successful with three different side-channel attacks on graphics cards via the WebGPU browser interface. The attacks…

A closer look at mechanochemistry
Ferdi Schüth and his team at the Max Planck Institut für Kohlenforschung in Mülheim/Germany have been studying the phenomena of mechanochemistry for several years. But what actually happens at the…

Severe Vulnerabilities Discovered in Software to Protect Internet Routing
A research team from the National Research Center for Applied Cybersecurity ATHENE led by Prof. Dr. Haya Schulmann has uncovered 18 vulnerabilities in crucial software components of Resource Public Key…