New technology enables 5-D imaging in live animals, humans
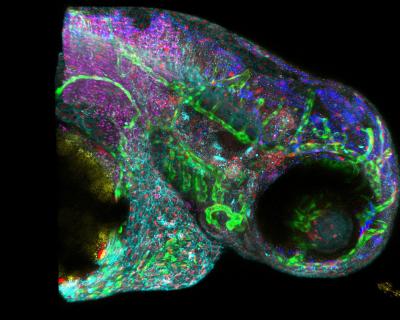
Novel image analysis technology enables new insights in complex systems biomolecules networks. By enhanced multi-color imaging scientists at USC's Translational Imaging Center access a new information dimension that reveals interactions between molecules. In the figure, a Zebrafish embryo with six different colors was analyzed by Francesco Cutrale and Scott E. Fraser at the University of Southern California's Translational Imaging Center. Credit: USC Translational Imaging Center
A new image analysis technique makes it easier for scientists to quickly find and track important biological molecules including tell-tale signs of disease.
Called “Hyper-Spectral Phasor” analysis, or HySP, it could even be useful for diagnosing and monitoring diseases by using cell phone images. It is much faster and far less expensive than current techniques.
Through the new imaging technology, researchers use fluorescent imaging to locate proteins and other molecules in cells and tissues. It works by tagging the molecules with dyes that glow under certain kinds of light — the same principle behind so-called “black light” images.
Fluorescent imaging can help scientists understand which molecules are produced in large amounts in cancer or other diseases, information that may be useful in diagnosis or in identifying possible targets for therapeutic drugs.
Looking at just one or two molecules in cell or tissue samples is fairly straightforward. Unfortunately, it doesn't provide a clear picture of how those molecules are behaving in the real world. For that, scientists need to expand their view.
“Biological research is moving toward complex systems that extend across multiple dimensions, the interaction of multiple elements over time,” said postdoctoral fellow Francesco Cutrale. He developed HySP with Scott Fraser, a USC Provost Professor of Biological Science who also holds the Elizabeth Garrett Chair in Convergent Bioscience.
The work was conducted at USC's Translational Imaging Center, a joint venture of USC Dornsife and USC Viterbi School of Engineering. It was published in the journal Nature Methods on Jan. 9.
“By looking at multiple targets, or watching targets move over time, we can get a much better view of what's actually happening within complex living systems,” Cutrale said. Currently, researchers must look at different labels separately, then apply complicated techniques to layer them together and figure out how they relate to one another, a time-consuming and expensive process, Cutrale said. HySP can look at many different molecules in one pass.
“Imagine looking at 18 targets,” Cutrale said. “We can do that all at once, rather than having to perform 18 separate experiments and try to combine them later.”
In addition, the algorithm effectively filters through interference to discern the true signal, even if that signal is extremely weak — very much like finding the proverbial needle in a haystack. Recent technology from NASA's Jet Propulsion Laboratory can also do this, but the equipment and process are both extremely expensive and time-consuming.
“HySP uses much less computing time, and we don't need the expensive imaging instrumentation,” Fraser said.
Cutrale and Fraser, along with researchers from Keck School of Medicine of USC, Caltech and the University of Cambridge in the United Kingdom, have used zebra fish to test and develop HySP. In this common laboratory model, the system works extremely well. But what about in people?
“In experimental models, we can use genetic manipulation to label molecules, but we can't do that with people,” said Fraser. “In people, we have to use the intrinsic signals of those molecules.”
Those inherent signals, the natural fluorescence from biomolecules, normally gets in the way of imaging, Fraser said. However, using this new computer algorithm that can effectively find weak signals in a cluttered background, the team can pinpoint their targets in the body.
The scientists hope to test the process in the next couple of years with the help of soldiers whose lungs have been damaged by chemicals and irritants they may have encountered in combat.
The researchers will extend a light-emitting probe into the soldiers' lungs that will record images of the fluorescence in the surrounding tissues. They will then use HySP to create what amounts to a fluorescent map and compare it with that of healthy lung tissue to see if they can discern the damage.
If they are successful, they hope to further develop the technology so it may one day help these soldiers and other lung patients receive more targeted treatment.
It might also be possible one day for clinicians to use HySP to analyze cell phone pictures of skin lesions to determine if they are at risk of being cancerous, said Fraser and Cutrale.
“We could determine if the lesions have changed color or shape over time,” Cutrale said. Clinicians could then examine the patient further to be certain of a diagnosis and respond appropriately.
Cutrale and Fraser see the technology as a giant leap forward for both research and medicine.
“Both scientists at the bench and scientists at the clinic will be able to perform their work faster and with greater confidence in the results,” Cutrale said. “Better, faster, cheaper. That's the payoff here.”
###
Other study co-authors were Vikas Trivedi of USC's Translational Imaging Center, Caltech and the University of Cambridge and USC's Le A. Trinh, Chi-Li Chiu, John M. Choi and Marcela S. Artiga.
This research was supported by grants from the Moore Foundation and the National Institutes of Health (R01 HD075605, R01 OD019037).
Media Contact
All latest news from the category: Information Technology
Here you can find a summary of innovations in the fields of information and data processing and up-to-date developments on IT equipment and hardware.
This area covers topics such as IT services, IT architectures, IT management and telecommunications.
Newest articles
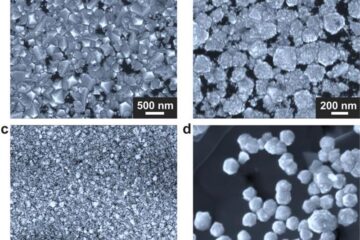

Making diamonds at ambient pressure
Scientists develop novel liquid metal alloy system to synthesize diamond under moderate conditions. Did you know that 99% of synthetic diamonds are currently produced using high-pressure and high-temperature (HPHT) methods?[2]…

Eruption of mega-magnetic star lights up nearby galaxy
Thanks to ESA satellites, an international team including UNIGE researchers has detected a giant eruption coming from a magnetar, an extremely magnetic neutron star. While ESA’s satellite INTEGRAL was observing…
Solving the riddle of the sphingolipids in coronary artery disease
Weill Cornell Medicine investigators have uncovered a way to unleash in blood vessels the protective effects of a type of fat-related molecule known as a sphingolipid, suggesting a promising new…