New software designed for rapid, automated identification of dendritic spines
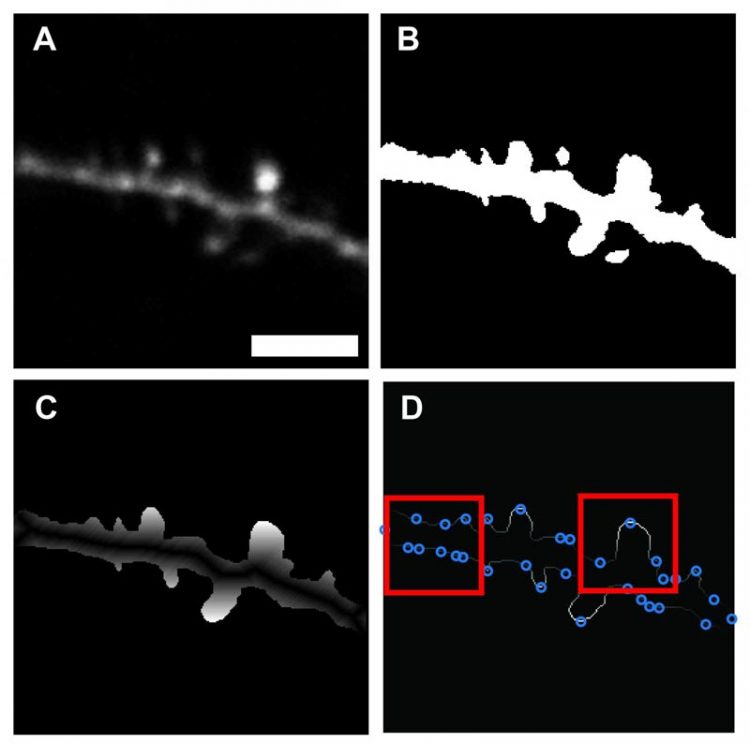
Once an image is loaded (A), pixels are converted to grayscale floating-point numbers ranging between 0 to 1. Highest probability background is identified using Otsu's method of globally thresholding. The resulting binary image (B) includes only regions of neuronal tissue. The location of each feature vector, as well as the individual values of perimeter distance features, were quantified based on a geodesic distance transform of the binary image of the dendrite (C), using the dendrite backbone as a seed location. Identification of potential spine locations by local maxima along perimeter (D). Credit: Max Planck Florida Institute for Neuroscience
Is it possible for microscopes to learn a bit about the brain? Even be taught by neuroscientists to reliably recognize parts of brain cells… all on their own? Though it may seem like something straight out of “The Jetsons”, a neuroscientist and software engineer in the lab of MPFI's Scientific Director, Dr. Ryohei Yasuda, Ph.D., is developing new software with the goal of vastly improving the daily life of a microscope user.
Combining a specialized algorithm, aptly named a neural network with a small amount of training, microscopes can now autonomously and efficiently identify small neuronal compartments called dendritic spines with over 90% accuracy. Just like Rosie the robotic maid, microscopes equipped with this Spine Identification software are helping scientists streamline their daily routine, bringing research one step into the future.
In recent years, there has been a rapid expansion and advancement in powerful imaging technologies, capable of taking a look inside the brain with unprecedented resolution and sensitivity. From imaging technology alone, neuroscientists have gained an immense wealth of new information about the brain. But despite many advances in implementation and capabilities, powerful microscopes (and the software running them) are still lacking when it comes to ease of use and overall user experience.
In a new study published in PLOS ONE, Dr. Michael Smirnov, Ph.D., has created an imaging software he hopes will craft a new type of experience for users.
Smirnov explains that “When engineers and scientists design cutting-edge microscopes, they usually focus on the actual physical components and design. They are mostly interested in what these imaging technologies can do, what boundaries they can break and how they perform. Far less attention is placed on how these complex technologies can be made accessible for the average user and really improve their workflow. Each time I write my software, I always think about the user first; how can I make a difference for the person using it, make their research a little easier.”
The Yasuda lab studies the complex process called synaptic plasticity, which is thought to be the cellular basis of learning and memory. When single dendritic spines are stimulated, hundreds of signaling molecules are mobilized to carry new information throughout the neuron. Members of the lab investigate this process in detail utilizing 2-photon microscopy, hoping to glean insights into how the interplay of these molecules translates into memory.
Live 2-photon imaging experiments can be an intensive endeavor. Scientists must patiently sift through a neuron's dendritic arbor, scanning hundreds of spines for suitable candidates to image. Experiments are frequently repeated to amass sufficient data and those that fail mid-way must be restarted. This often unspoken aspect can quickly turn prolonged imaging into a tedious and time consuming chore, ultimately slowing scientific progress.
Dr. Smirnov's software aims to rescue neuroscientists from the humdrum of spine imaging. Weaving in an element of machine learning, the algorithm can be taught how to differentiate between dendrite backbone and dendritic spines after being fed a training data set of previously identified spines.
Once the training period finishes, the software is capable of automatedly scanning through an image and demarcating spines it comes across with high precision. Unlike previously developed programs which can be computing intensive or optimized only for post imaging analysis, Smirnov's software is fast, scalable and compatible with most live imaging setups and post analysis applications.
“We incorporated a machine learning approach, because we wanted our software to be flexible, adaptable and as easy to use as possible.” Says Smirnov “The program only requires the user to input an image and specify the scale. After that, the program does the rest.”
Smirnov describes that by automating the process of spine identification, the software has the potential to dramatically increase experiment workflow, shaving off hours of time. In addition, he has also made his program's code readily available to the broader neuroscience community. A move he hopes will engage the community of coders working in the field and give them the ability to freely improve upon and customize the software, making it useful for wider applications.
Dr. Smirnov notes that Scientists at MPFI are not only committed to conducting high caliber basic neuroscience research, but also look for ways to continuously improve the research process; fast tracking new discoveries and positively impacting scientists' daily lives.
Media Contact
All latest news from the category: Information Technology
Here you can find a summary of innovations in the fields of information and data processing and up-to-date developments on IT equipment and hardware.
This area covers topics such as IT services, IT architectures, IT management and telecommunications.
Newest articles

Superradiant atoms could push the boundaries of how precisely time can be measured
Superradiant atoms can help us measure time more precisely than ever. In a new study, researchers from the University of Copenhagen present a new method for measuring the time interval,…

Ion thermoelectric conversion devices for near room temperature
The electrode sheet of the thermoelectric device consists of ionic hydrogel, which is sandwiched between the electrodes to form, and the Prussian blue on the electrode undergoes a redox reaction…

Zap Energy achieves 37-million-degree temperatures in a compact device
New publication reports record electron temperatures for a small-scale, sheared-flow-stabilized Z-pinch fusion device. In the nine decades since humans first produced fusion reactions, only a few fusion technologies have demonstrated…