AI reveals nature of RNA-protein interactions
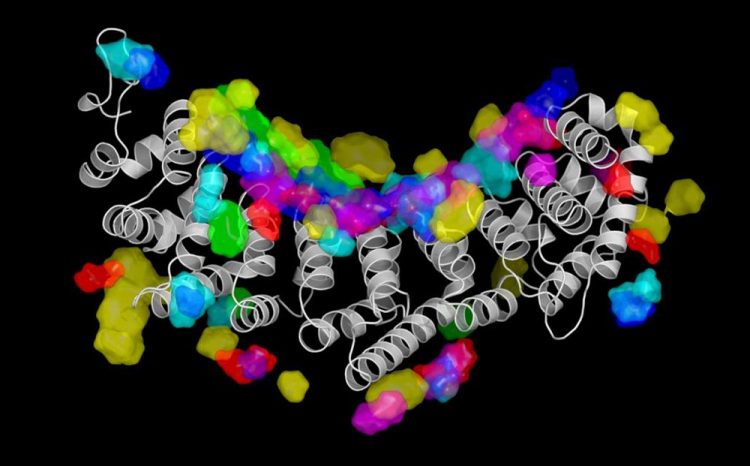
This illustration depicts the RNA interaction surface of Fem-3-binding-factor 2 as predicted by NucleicNet. Credit: 2019 KAUST
The software, called NucleicNet, outperforms other algorithmic models of its kind and provides additional biological insights that could aid in drug design and development.
“RNA binding is a fundamental feature of many proteins,” says Jordy Homing Lam, a former research associate at KAUST and co-first author of the study.
“Our structure-based computational framework can reveal the detailed RNA-binding properties of these proteins, which is important for characterizing the pathology of many diseases.”
Proteins routinely interface with RNA molecules as a way to control the processing and transporting of gene transcripts–and when these interactions go awry, information flow inside the cell is disrupted and disorders can arise, including cancer and neurodegenerative disease.
To better understand which parts of an RNA molecule tend to bind on different surface points of a protein, Lam and his colleagues turned to deep learning, a type of AI.
Working in the laboratory of KAUST Professor Xin Gao in the Computational Bioscience Research Center, Lam and Ph.D. student Yu Li, taught NucleicNet to automatically learn the structural features that underpin interactions between proteins and RNA.
They trained the algorithm using three-dimensional structural data from 158 different protein-RNA complexes available on a public database. Pitting NucleicNet against other predictive models–all of which rely on sequence inputs rather than structural information–the KAUST team showed that the tool could most accurately detect which sites on a protein surface bound to RNA molecules or not.
What's more, unlike any other model, NucleicNet could predict which aspects of the RNA molecule were doing the binding, be it part of the sugar-phosphate backbone or one of the four letters of the genetic alphabet.
In collaboration with researchers in China and the United States, Lam, Li and Gao validated their algorithm on a diverse set of RNA-binding proteins, including proteins implicated in gum cancer and amyotrophic lateral sclerosis, to show that the interactions deduced by NucleicNet closely matched those revealed by experimental techniques. They reported the findings in Nature Communications.
“Structure-based features were little considered by other computational frameworks,” says Lam. “We have harnessed the power of deep learning to infer those subtle interactions.”
NucleicNet is openly available for researchers who want to predict RNA-binding sites and binding preference for any protein of interest. The software can be accessed at http://www.
Media Contact
More Information:
http://dx.doi.org/10.1038/s41467-019-12920-0All latest news from the category: Information Technology
Here you can find a summary of innovations in the fields of information and data processing and up-to-date developments on IT equipment and hardware.
This area covers topics such as IT services, IT architectures, IT management and telecommunications.
Newest articles

Bringing bio-inspired robots to life
Nebraska researcher Eric Markvicka gets NSF CAREER Award to pursue manufacture of novel materials for soft robotics and stretchable electronics. Engineers are increasingly eager to develop robots that mimic the…

Bella moths use poison to attract mates
Scientists are closer to finding out how. Pyrrolizidine alkaloids are as bitter and toxic as they are hard to pronounce. They’re produced by several different types of plants and are…

AI tool creates ‘synthetic’ images of cells
…for enhanced microscopy analysis. Observing individual cells through microscopes can reveal a range of important cell biological phenomena that frequently play a role in human diseases, but the process of…